Lesson type
Difficulty level
Course:
This video will document the process of launching a Jupyter Notebook for group-level analyses directly from brainlife.
Difficulty level: Intermediate
Duration: 0:53
Speaker: :
Course:
This lesson is a general overview of overarching concepts in neuroinformatics research, with a particular focus on clinical approaches to defining, measuring, studying, diagnosing, and treating various brain disorders. Also described are the complex, multi-level nature of brain disorders and the data associated with them, from genes and individual cells up to cortical microcircuits and whole-brain network dynamics. Given the heterogeneity of brain disorders and their underlying mechanisms, this lesson lays out a case for multiscale neuroscience data integration.
Difficulty level: Intermediate
Duration: 1:09:33
Speaker: : Sean Hill
This is a continuation of the talk on the cellular mechanisms of neuronal communication, this time at the level of brain microcircuits and associated global signals like those measureable by electroencephalography (EEG). This lecture also discusses EEG biomarkers in mental health disorders, and how those cortical signatures may be simulated digitally.
Difficulty level: Intermediate
Duration: 1:11:04
Speaker: : Etay Hay
This lecture aims to help researchers, students, and health care professionals understand the place for neuroinformatics in the patient journey using the exemplar of an epilepsy patient.
Difficulty level: Intermediate
Duration: 1:32:53
Speaker: : Randy Gollub & Prantik Kundu
The lesson introduces the Brain Imaging Data Structure (BIDS), the community standard for organizing, curating, and sharing neuroimaging and associated data. The session focuses on understanding the BIDS framework, learning its data structure and validation processes.
Difficulty level: Intermediate
Duration: 38:52
Speaker: : Cyril Pernet
This session moves from BIDS basics into analysis workflows, focusing on how to turn raw, BIDS-organized data into derivatives using BIDS Apps and containers for reproducible processing. It compares end-to-end pipelines across fMRI and PET (and notes EEG/MEG), explains typical preprocessing choices, and shows how standardized inputs plus containerized tools (Docker/AppTainer) yield consistent, auditable outputs.
Difficulty level: Intermediate
Duration: 56:03
Speaker: : Martin Nørgaard
The session explains GDPR rules around data sharing for research in Europe, the distinction between law and ethics, and introduces practical solutions for securely sharing sensitive datasets. Researchers have more flexibility than commonly assumed: scientific research is considered a public interest task, so explicit consent for data sharing isn’t legally required, though transparency and informing participants remain ethically important. The talk also introduces publicneuro.eu, a controlled-access platform that enables sharing neuroimaging datasets with open metadata, DOIs, and customizable access restrictions while ensuring GDPR compliance.
Difficulty level: Intermediate
Duration: 31:12
Speaker: : Cyril Pernet
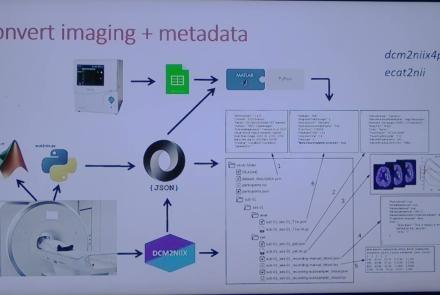
This session introduces the PET-to-BIDS (PET2BIDS) library, a toolkit designed to simplify the conversion and preparation of PET imaging datasets into BIDS-compliant formats. It supports multiple data types and formats (e.g., DICOM, ECAT7+, nifti, JSON), integrates seamlessly with Excel-based metadata, and provides automated routines for metadata updates, blood data conversion, and JSON synchronization. PET2BIDS improves human readability by mapping complex reconstruction names into standardized, descriptive labels and offers extensive documentation, examples, and video tutorials to make adoption easier for researchers.
Difficulty level: Intermediate
Duration: 9:23
Speaker: : Cyril Pernet
This session introduces the PET-to-BIDS (PET2BIDS) library, a toolkit designed to simplify the conversion and preparation of PET imaging datasets into BIDS-compliant formats. It supports multiple data types and formats (e.g., DICOM, ECAT7+, nifti, JSON), integrates seamlessly with Excel-based metadata, and provides automated routines for metadata updates, blood data conversion, and JSON synchronization. PET2BIDS improves human readability by mapping complex reconstruction names into standardized, descriptive labels and offers extensive documentation, examples, and video tutorials to make adoption easier for researchers.
Difficulty level: Intermediate
Duration: 41:04
Speaker: : Martin Nørgaard
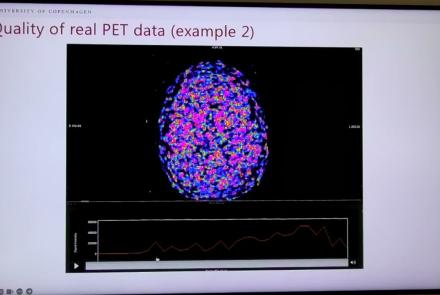
This session dives into practical PET tooling on BIDS data—showing how to run motion correction, register PET↔MRI, extract time–activity curves, and generate standardized PET-BIDS derivatives with clear QC reports. It introduces modular BIDS Apps (head-motion correction, TAC extraction), a full pipeline (PETPrep), and a PET/MRI defacer, with guidance on parameters, outputs, provenance, and why Dockerized containers are the reliable way to run them at scale.
Difficulty level: Intermediate
Duration: 1:05:38
Speaker: : Martin Nørgaard
This session introduces two PET quantification tools—bloodstream for processing arterial blood data and kinfitr for kinetic modeling and quantification—built to work with BIDS/BIDS-derivatives and containers. Bloodstream fuses autosampler and manual measurements (whole blood, plasma, parent fraction) using interpolation or fitted models (incl. hierarchical GAMs) to produce a clean arterial input function (AIF) and whole-blood curves with rich QC reports ready. TAC data (e.g., from PETPrep) and blood (e.g., from bloodstream) can be ingested using kinfitr to run reproducible, GUI-driven analyses: define combined ROIs, calculate weighting factors, estimate blood–tissue delay, choose and chain models (e.g., 2TCM → 1TCM with parameter inheritance), and export parameters/diagnostics. Both are available as Docker apps; workflows emphasize configuration files, reports, and standard outputs to support transparency and reuse.
Difficulty level: Intermediate
Duration: 1:20:56
Speaker: : Granville Matheson
This lecture provides an introduction to entropy in general, and multi-scale entropy (MSE) in particular, highlighting the potential clinical applications of the latter.
Difficulty level: Intermediate
Duration: 39:05
Speaker: : Jil Meier
This lecture provides an general introduction to epilepsy, as well as why and how TVB can prove useful in building and testing epileptic models.
Difficulty level: Intermediate
Duration: 37:12
Speaker: : Julie Courtiol
Course:
This lecture covers the rationale for developing the DAQCORD, a framework for the design, documentation, and reporting of data curation methods in order to advance the scientific rigour, reproducibility, and analysis of data.
Difficulty level: Intermediate
Duration: 17:08
Speaker: : Ari Ercole
This lecture focuses on ontologies for clinical neurosciences.
Difficulty level: Intermediate
Duration: 21:54
Speaker: : Martin Hofmann-Apitius
This talks presents an overview of the potential for data federation in stroke research.
Difficulty level: Intermediate
Duration: 21:37
Speaker: : Maurizio A. Leone
This lecture explains the need for data federation in medicine and how it can be achieved.
Difficulty level: Intermediate
Duration: 27:09
Speaker: : Philippe Ryvlin
In this session the Medical Informatics Platform (MIP) federated analytics is presented. The current and future analytical tools implemented in the MIP will be detailed along with the constructs, tools, processes, and restrictions that formulate the solution provided. MIP is a platform providing advanced federated analytics for diagnosis and research in clinical neuroscience research. It is targeting clinicians, clinical scientists and clinical data scientists. It is designed to help adopt advanced analytics, explore harmonized medical data of neuroimaging, neurophysiological and medical records as well as research cohort datasets, without transferring original clinical data. It can be perceived as a virtual database that seamlessly presents aggregated data from distributed sources, provides access and analyze imaging and clinical data, securely stored in hospitals, research archives and public databases. It leverages and re-uses decentralized patient data and research cohort datasets, without transferring original data. Integrated statistical analysis tools and machine learning algorithms are exposed over harmonized, federated medical data.
Difficulty level: Intermediate
Duration: 15:05
Speaker: : Giorgos Papanikos
The Medical Informatics Platform (MIP) is a platform providing federated analytics for diagnosis and research in clinical neuroscience research. The federated analytics is possible thanks to a distributed engine that executes computations and transfers information between the members of the federation (hospital nodes). In this talk the speaker will describe the process of designing and implementing new analytical tools, i.e. statistical and machine learning algorithms. Mr. Sakellariou will further describe the environment in which these federated algorithms run, the challenges and the available tools, the principles that guide its design and the followed general methodology for each new algorithm. One of the most important challenges which are faced is to design these tools in a way that does not compromise the privacy of the clinical data involved. The speaker will show how to address the main questions when designing such algorithms: how to decompose and distribute the computations and what kind of information to exchange between nodes, in order to comply with the privacy constraint mentioned above. Finally, also the subject of validating these federated algorithms will be briefly touched.
Difficulty level: Intermediate
Duration: 20:26
Speaker: : Jason Skellariou
The Medical Informatics Platform (MIP) Dementia had been installed in several memory clinics across Europe allowing them to federate their real-world databases. Research open access databases had also been integrated such as ADNI (Alzheimer’s Dementia Neuroimaging Initiative), reaching a cumulative case load of more than 5,000 patients (major cognitive disorder due to Alzheimer’s disease, other major cognitive disorder, minor cognitive disorder, controls). The statistic and machine learning tools implemented in the MIP allowed researchers to conduct easily federated analyses among Italian memory clinics (Redolfi et al. 2020) and also across borders between the French (Lille), the Swiss (Lausanne) and the Italian (Brescia) datasets.
Difficulty level: Intermediate
Duration: 16:44
Speaker: : Mélanie Leroy
Topics
- Artificial Intelligence (1)
- (-) Notebooks (1)
- Provenance (1)
- DANDI archive (1)
- EBRAINS RI (6)
- Animal models (2)
- Brain-hardware interfaces (1)
- Clinical neuroscience (23)
- General neuroscience
(17)
- General neuroinformatics
(1)
- Computational neuroscience (81)
- Statistics (5)
- Computer Science (5)
- Genomics (8)
- Data science
(10)
- Open science (5)
- Project management (1)
- Education (1)
- Neuroethics (5)