Course:
This lecture will provide an overview of the INCF Training Suite, a collection of tools that embraces the FAIR principles developed by members of the INCF Community. This will include an overview of TrainingSpace, Neurostars, and KnowledgeSpace.
Difficulty level: Beginner
Duration: 09:50
Speaker: : Mathew Abrams
Course:
This session will include presentations of infrastructure that embrace the FAIR principles developed by members of the INCF Community. This lecture provides an overview and demo of the Canadian Open Neuroscience Platform (CONP).
Difficulty level: Beginner
Duration: 14:02
Speaker: : Tristan Glatard & Samir Das
This lecture contains an overview of the China-Cuba-Canada neuroinformatics ecosystem for Quantitative Tomographic EEG Analysis (qEEGt).
Difficulty level: Beginner
Duration: 12:56
Speaker: : Pedro Valdes-Sosa
Course:
This session provides users with an introduction to tools and resources that facilitate the implementation of FAIR in their research.
Difficulty level: Beginner
Duration: 38:36
This lesson provides an overview of self-supervision as it relates to neural data tasks and the Mine Your Own vieW (MYOW) approach.
Difficulty level: Beginner
Duration: 25:50
Speaker: : Eva Dyer
In this talk, you will learn how brainlife.io works, and how it can be applied to neuroscience data.
Difficulty level: Beginner
Duration: 10:14
Speaker: : Franco Pestilli
This lecture covers the IBI Data Standards and Sharing Working Group, including its history, aims, and projects.
Difficulty level: Beginner
Duration: 3:58
Speaker: : Kenji Doya
This session covers the framework of the International Brain Lab (IBL) and the data architecture used for this project.
Difficulty level: Beginner
Duration: 23:37
Speaker: : Kenneth Harris
Course:
This session will include presentations of infrastructure that embrace the FAIR principles developed by members of the INCF Community.
This lecture provides an overview of The Virtual Brain Simulation Platform.
Difficulty level: Beginner
Duration: 9:36
Speaker: : Petra Ritter
This lesson provides an overview of the current status in the field of neuroscientific ontologies, presenting examples of data organization and standards, particularly from neuroimaging and electrophysiology.
Difficulty level: Intermediate
Duration: 33:41
Speaker: : Yaroslav O. Halchenko
Hierarchical Event Descriptors (HED) fill a major gap in the neuroinformatics standards toolkit, namely the specification of the nature(s) of events and time-limited conditions recorded as having occurred during time series recordings (EEG, MEG, iEEG, fMRI, etc.). Here, the HED Working Group presents an online INCF workshop on the need for, structure of, tools for, and use of HED annotation to prepare neuroimaging time series data for storing, sharing, and advanced analysis.
Difficulty level: Beginner
Duration: 03:37:42
Speaker: :
In this lesson, attendees will learn about the data structure standards, specifically the Brain Imaging Data Structure (BIDS), an INCF-endorsed standard for organizing, annotating, and describing data collected during neuroimaging experiments.
Difficulty level: Beginner
Duration: 21:56
Speaker: : Michael Schirner
This lightning talk describes an automated pipline for positron emission tomography (PET) data.
Difficulty level: Intermediate
Duration: 7:27
Speaker: : Soodeh Moallemian
Course:
This module covers many of the types of non-invasive neurotech and neuroimaging devices including electroencephalography (EEG), electromyography (EMG), electroneurography (ENG), magnetoencephalography (MEG), and more.
Difficulty level: Beginner
Duration: 13:36
Speaker: : Harrison Canning
This lesson contains practical exercises which accompanies the first few lessons of the Neuroscience for Machine Learners (Neuro4ML) course.
Difficulty level: Intermediate
Duration: 5:58
Speaker: : Dan Goodman
This video briefly goes over the exercises accompanying Week 6 of the Neuroscience for Machine Learners (Neuro4ML) course, Understanding Neural Networks.
Difficulty level: Intermediate
Duration: 2:43
Speaker: : Marcus Ghosh
Course:
This lecture covers the description and characterization of an input-output relationship in a information-theoretic context.
Difficulty level: Beginner
Duration: 1:35:33
Speaker: : Jonathan D. Victor
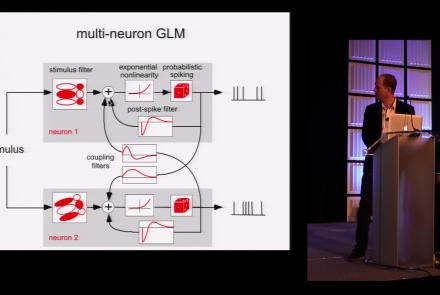
This lesson is part 1 of 2 of a tutorial on statistical models for neural data.
Difficulty level: Beginner
Duration: 1:45:48
Speaker: : Jonathan Pillow
This lesson is part 2 of 2 of a tutorial on statistical models for neural data.
Difficulty level: Beginner
Duration: 1:50:31
Speaker: : Jonathan Pillow
Course:
This lesson provides an introduction to modeling single neurons, as well as stability analysis of neural models.
Difficulty level: Intermediate
Duration: 1:26:06
Speaker: : Bard Ermentrout
Topics
- Artificial Intelligence (7)
- Philosophy of Science (5)
- Provenance (3)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (8)
- (-) Assembly 2021 (29)
- Brain-hardware interfaces (14)
- Clinical neuroscience (40)
- International Brain Initiative (2)
- Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(62)
- Neuroscience (11)
- Cognitive Science (7)
- Cell signaling (6)
- Brain networks (11)
- (-) Glia (1)
- (-) Electrophysiology (41)
- Learning and memory (5)
- (-) Neuroanatomy (24)
- Neurobiology (16)
- Neurodegeneration (1)
- Neuroimmunology (1)
- Neural networks (15)
- Neurophysiology (27)
- Neuropharmacology (2)
- Neuronal plasticity (16)
- Synaptic plasticity (4)
- Visual system (12)
- Phenome (1)
- General neuroinformatics
(27)
- Computational neuroscience (275)
- Statistics (7)
- Computer Science (20)
- (-) Genomics (32)
- Data science
(33)
- Open science (57)
- Project management (8)
- Education (4)
- Publishing (4)
- Neuroethics (40)