Course:
This tutorial is part 1 of 2. It aims to provide viewers with an understanding of the fundamentals of R tool. Note: parts 1 and 2 of this tutorial are part of the same YouTube video; part 1 ends at 17:42.
Difficulty level: Beginner
Duration: 17:42
Speaker: : Edureka
This lesson introduces the practical usage of The Virtual Brain (TVB) in its graphical user interface and via python scripts. In the graphical user interface, you are guided through its data repository, simulator, phase plane exploration tool, connectivity editor, stimulus generator, and the provided analyses. The implemented iPython notebooks of TVB are presented, and since they are public, can be used for further exploration of TVB.
Difficulty level: Beginner
Duration: 1:12:24
Speaker: : Paul Triebkorn
Course:
This lesson provides a comprehensive introduction to the command line and 50 popular Linux commands. This is a long introduction (nearly 5 hours), but well worth it if you are going to spend a good part of your career working from a terminal, which is likely if you are interested in flexibility, power, and reproducibility in neuroscience research. This lesson is courtesy of freeCodeCamp.
Difficulty level: Beginner
Duration: 5:00:16
Speaker: : Colt Steele
Research Resource Identifiers (RRIDs) are ID numbers assigned to help researchers cite key resources (e.g., antibodies, model organisms, and software projects) in biomedical literature to improve the transparency of research methods.
Difficulty level: Beginner
Duration: 1:01:36
Speaker: : Maryann Martone
Course:
This lesson introduces the EEGLAB toolbox, as well as motivations for its use.
Difficulty level: Beginner
Duration: 15:32
Speaker: : Arnaud Delorme
Course:
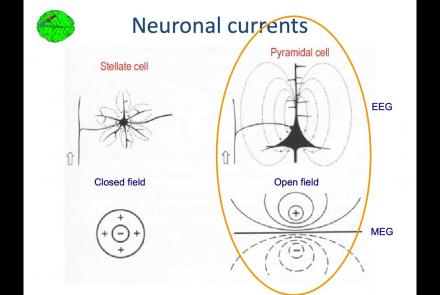
In this lesson, you will learn about the biological activity which generates and is measured by the EEG signal.
Difficulty level: Beginner
Duration: 6:53
Speaker: : Arnaud Delorme
Course:
This lesson goes over the characteristics of EEG signals when analyzed in source space (as opposed to sensor space).
Difficulty level: Beginner
Duration: 10:56
Speaker: : Arnaud Delorme
Course:
This lesson describes the development of EEGLAB as well as to what extent it is used by the research community.
Difficulty level: Beginner
Duration: 6:06
Speaker: : Arnaud Delorme
Course:
This lesson provides instruction as to how to build a processing pipeline in EEGLAB for a single participant.
Difficulty level: Beginner
Duration: 9:20
Speaker: :
Course:
Whereas the previous lesson of this course outlined how to build a processing pipeline for a single participant, this lesson discusses analysis pipelines for multiple participants simultaneously.
Difficulty level: Beginner
Duration: 10:55
Speaker: : Arnaud Delorme
Course:
In addition to outlining the motivations behind preprocessing EEG data in general, this lesson covers the first step in preprocessing data with EEGLAB, importing raw data.
Difficulty level: Beginner
Duration: 8:30
Speaker: : Arnaud Delorme
Course:
Continuing along the EEGLAB preprocessing pipeline, this tutorial walks users through how to import data events as well as EEG channel locations.
Difficulty level: Beginner
Duration: 11:53
Speaker: : Arnaud Delorme
Course:
This tutorial demonstrates how to re-reference and resample raw data in EEGLAB, why such steps are important or useful in the preprocessing pipeline, and how choices made at this step may affect subsequent analyses.
Difficulty level: Beginner
Duration: 11:48
Speaker: : Arnaud Delorme
Course:
This tutorial instructs users how to visually inspect partially pre-processed neuroimaging data in EEGLAB, specifically how to use the data browser to investigate specific channels, epochs, or events for removable artifacts, biological (e.g., eye blinks, muscle movements, heartbeat) or otherwise (e.g., corrupt channel, line noise).
Difficulty level: Beginner
Duration: 5:08
Speaker: : Arnaud Delorme
Course:
This tutorial provides instruction on how to use EEGLAB to further preprocess EEG datasets by identifying and discarding bad channels which, if left unaddressed, can corrupt and confound subsequent analysis steps.
Difficulty level: Beginner
Duration: 13:01
Speaker: : Arnaud Delorme
Course:
Users following this tutorial will learn how to identify and discard bad EEG data segments using the MATLAB toolbox EEGLAB.
Difficulty level: Beginner
Duration: 11:25
Speaker: : Arnaud Delorme
Course:
This module covers many of the types of non-invasive neurotech and neuroimaging devices including electroencephalography (EEG), electromyography (EMG), electroneurography (ENG), magnetoencephalography (MEG), and more.
Difficulty level: Beginner
Duration: 13:36
Speaker: : Harrison Canning
Course:
In this lesson, while learning about the need for increased large-scale collaborative science that is transparent in nature, users also are given a tutorial on using Synapse for facilitating reusable and reproducible research.
Difficulty level: Beginner
Duration: 1:15:12
Speaker: : Abhi Pratap
This lecture covers advanced concept of energy based models. The lecture is a part of the Advanced energy based models modules of the the Deep Learning Course at NYU's Center for Data Science. Prerequisites for this course include: Energy-Based Models I, Energy-Based Models II, Energy-Based Models III, and an Introduction to Data Science or a Graduate Level Machine Learning course.
Difficulty level: Beginner
Duration: 56:41
Speaker: : Alfredo Canziani
Course:
As a part of NeuroHackademy 2021, Noah Benson gives an introduction to Pytorch, one of the two most common software packages for deep learning applications to the neurosciences.
Difficulty level: Beginner
Duration: 00:50:40
Speaker: :
Topics
- Electroencephalography (EEG) (14)
- Provenance (2)
- (-) Standards and Best Practices (1)
- Brain Medicine (1)
- Artificial Intelligence (1)
- Notebooks (1)
- Event related potential (ERP) (13)
- Digital brain atlasing (3)
- Neuroimaging (16)
- Epilepsy (1)
- Optogenetics (1)
- Neurodevelopmental disorders (1)
- Standards and best practices (1)
- Tools (13)
- Psychology (1)
- Stroke (1)
- Workflows (1)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Assembly 2021 (1)
- Brain-hardware interfaces (12)
- Repositories and science gateways (1)
- Resources (1)
- General neuroscience (5)
- Phenome (1)
- Computational neuroscience (62)
- Statistics (2)
- (-) Computer Science (3)
- (-) Genomics (23)
- Data science (12)
- Open science (14)
- Education (1)
- Neuroethics (1)