This is a tutorial on designing a Bayesian inference model to map belief trajectories, with emphasis on gaining familiarity with Hierarchical Gaussian Filters (HGFs).
This lesson corresponds to slides 65-90 of the PDF below.
Difficulty level: Intermediate
Duration: 1:15:04
Speaker: : Daniel Hauke
Similarity Network Fusion (SNF) is a computational method for data integration across various kinds of measurements, aimed at taking advantage of the common as well as complementary information in different data types. This workshop walks participants through running SNF on EEG and genomic data using RStudio.
Difficulty level: Intermediate
Duration: 1:21:38
Speaker: : Dan Felsky
This lecture aims to help researchers, students, and health care professionals understand the place for neuroinformatics in the patient journey using the exemplar of an epilepsy patient.
Difficulty level: Intermediate
Duration: 1:32:53
Speaker: : Randy Gollub & Prantik Kundu
This lesson continues from part one of the lecture Ontologies, Databases, and Standards, diving deeper into a description of ontologies and knowledg graphs.
Difficulty level: Intermediate
Duration: 50:18
Speaker: : Jeff Grethe
This lecture describes how to build research workflows, including a demonstrate using DataJoint Elements to build data pipelines.
Difficulty level: Intermediate
Duration: 47:00
Speaker: : Dimitri Yatsenko
This lesson introduces some practical exercises which accompany the Synapses and Networks portion of this Neuroscience for Machine Learners course.
Difficulty level: Intermediate
Duration: 3:51
Speaker: : Dan Goodman
This lesson characterizes different types of learning in a neuroscientific and cellular context, and various models employed by researchers to investigate the mechanisms involved.
Difficulty level: Intermediate
Duration: 3:54
Speaker: : Dan Goodman
In this lesson, you will learn about different approaches to modeling learning in neural networks, particularly focusing on system parameters such as firing rates and synaptic weights impact a network.
Difficulty level: Intermediate
Duration: 9:40
Speaker: : Dan Goodman
This lesson explores how researchers try to understand neural networks, particularly in the case of observing neural activity.
Difficulty level: Intermediate
Duration: 8:20
Speaker: : Marcus Ghosh
As the previous lesson of this course described how researchers acquire neural data, this lesson will discuss how to go about interpreting and analysing the data.
Difficulty level: Intermediate
Duration: 9:24
Speaker: : Marcus Ghosh
In this lesson you will learn about the motivation behind manipulating neural activity, and what forms that may take in various experimental designs.
Difficulty level: Intermediate
Duration: 8:42
Speaker: : Marcus Ghosh
In this lesson, you will learn about one particular aspect of decision making: reaction times. In other words, how long does it take to take a decision based on a stream of information arriving continuously over time?
Difficulty level: Intermediate
Duration: 6:01
Speaker: : Dan Goodman
In this lesson, you will hear about some of the open issues in the field of neuroscience, as well as a discussion about whether neuroscience works, and how can we know?
Difficulty level: Intermediate
Duration: 6:54
Speaker: : Marcus Ghosh
This lesson discusses a gripping neuroscientific question: why have neurons developed the discrete action potential, or spike, as a principle method of communication?
Difficulty level: Intermediate
Duration: 9:34
Speaker: : Dan Goodman
This tutorial provides instruction on how to simulate brain tumors with TVB (reproducing publication: Marinazzo et al. 2020 Neuroimage). This tutorial comprises a didactic video, jupyter notebooks, and full data set for the construction of virtual brains from patients and health controls.
Difficulty level: Intermediate
Duration: 10:01
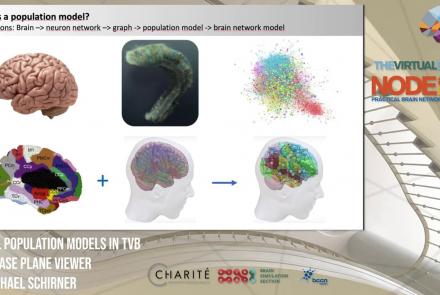
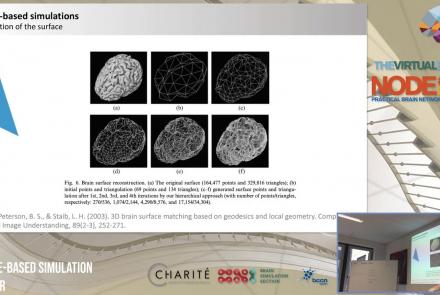
This lesson introduces population models and the phase plane, and is part of the The Virtual Brain (TVB) Node 10 Series, a 4-day workshop dedicated to learning about the full brain simulation platform TVB, as well as brain imaging, brain simulation, personalised brain models, and TVB use cases.
Difficulty level: Intermediate
Duration: 1:10:41
Speaker: : Michael Schirner
This lesson introduces TVB-multi-scale extensions and other TVB tools which facilitate modeling and analyses of multi-scale data.
Difficulty level: Intermediate
Duration: 36:10
Speaker: : Dionysios Perdikis
This lecture delves into cortical (i.e., surface-based) brain simulations, as well as subcortical (i.e., deep brain) stimulations, covering the definitions, motivations, and implementations of both.
Difficulty level: Intermediate
Duration: 39:05
Speaker: : Jil Meier
This lecture provides an introduction to entropy in general, and multi-scale entropy (MSE) in particular, highlighting the potential clinical applications of the latter.
Difficulty level: Intermediate
Duration: 39:05
Speaker: : Jil Meier
This lecture gives an overview of how to prepare and preprocess neuroimaging (EEG/MEG) data for use in TVB.
Difficulty level: Intermediate
Duration: 1:40:52
Speaker: : Paul Triebkorn
Topics
- Artificial Intelligence (1)
- Provenance (1)
- EBRAINS RI (6)
- Animal models (1)
- Brain-hardware interfaces (1)
- Clinical neuroscience (23)
- (-) General neuroscience (13)
- General neuroinformatics (1)
- Computational neuroscience (42)
- Statistics (5)
- (-) Computer Science (2)
- (-) Genomics (7)
- Data science
(8)
- Open science (3)
- Project management (1)
- Neuroethics (5)