Lesson type
Difficulty level
This lesson continues with the second workshop on reproducible science, focusing on additional open source tools for researchers and data scientists, such as the R programming language for data science, as well as associated tools like RStudio and R Markdown. Additionally, users are introduced to Python and iPython notebooks, Google Colab, and are given hands-on tutorials on how to create a Binder environment, as well as various containers in Docker and Singularity.
Difficulty level: Beginner
Duration: 1:16:04
Speaker: : Erin Dickie and Sejal Patel
This talk describes the relevance and power of using brain atlases as part of one's data integration pipeline.
Difficulty level: Beginner
Duration: 15:37
Speaker: : Timo Dickscheid
In this lesson, you will learn how to understand data management plans and why data sharing is important.
Difficulty level: Beginner
Duration: 44:24
Speaker: : Jenny Muilenburg
Course:
This quick visual walkthrough presents the steps required in uploading data into a brainlife project using the graphical user interface (GUI).
Difficulty level: Beginner
Duration: 2:00
Speaker: :
Course:
This short walkthrough documents the steps needed to find a dataset in OpenNeuro, a free and open platform for sharing MRI, MEG, EEG, iEEG, ECoG, ASL, and PET data, and import it directly to a brainlife project.
Difficulty level: Beginner
Duration: 0:35
Speaker: :
Course:
This lesson describes and shows four different ways one may upload their data to brainlife.io.
Difficulty level: Beginner
Duration: 6:35
Speaker: :
Course:
This lecture covers why data sharing and other collaborative practices are important, how these practices are developed, and the challenges involved in their development and implementation.
Difficulty level: Beginner
Duration: 11:41
Speaker: : Joost Wagenaar
This lecture discusses the FAIR principles as they apply to electrophysiology data and metadata, the building blocks for community tools and standards, platforms and grassroots initiatives, and the challenges therein.
Difficulty level: Beginner
Duration: 8:11
Speaker: : Thomas Wachtler
This lecture contains an overview of electrophysiology data reuse within the EBRAINS ecosystem.
Difficulty level: Beginner
Duration: 15:57
Speaker: : Andrew Davison
This lecture contains an overview of the Distributed Archives for Neurophysiology Data Integration (DANDI) archive, its ties to FAIR and open-source, integrations with other programs, and upcoming features.
Difficulty level: Beginner
Duration: 13:34
Speaker: : Yaroslav O. Halchenko
This lesson provides a short overview of the main features of the Canadian Open Neuroscience Platform (CONP) Portal, a web interface that facilitates open science for the neuroscience community by simplifying global access to and sharing of datasets and tools. The Portal internalizes the typical cycle of a research project, beginning with data acquisition, followed by data processing with published tools, and ultimately the publication of results with a link to the original dataset.
Difficulty level: Beginner
Duration: 14:03
Speaker: : Samir Das, Tristan Glatard
This lesson contains practical exercises which accompanies the first few lessons of the Neuroscience for Machine Learners (Neuro4ML) course.
Difficulty level: Intermediate
Duration: 5:58
Speaker: : Dan Goodman
This video briefly goes over the exercises accompanying Week 6 of the Neuroscience for Machine Learners (Neuro4ML) course, Understanding Neural Networks.
Difficulty level: Intermediate
Duration: 2:43
Speaker: : Marcus Ghosh
Course:
This lecture covers the description and characterization of an input-output relationship in a information-theoretic context.
Difficulty level: Beginner
Duration: 1:35:33
Speaker: : Jonathan D. Victor
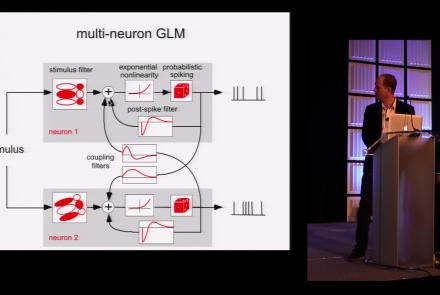
This lesson is part 1 of 2 of a tutorial on statistical models for neural data.
Difficulty level: Beginner
Duration: 1:45:48
Speaker: : Jonathan Pillow
This lesson is part 2 of 2 of a tutorial on statistical models for neural data.
Difficulty level: Beginner
Duration: 1:50:31
Speaker: : Jonathan Pillow
Course:
This lesson provides an introduction to modeling single neurons, as well as stability analysis of neural models.
Difficulty level: Intermediate
Duration: 1:26:06
Speaker: : Bard Ermentrout
Course:
This lesson continues a thorough description of the concepts, theories, and methods involved in the modeling of single neurons.
Difficulty level: Intermediate
Duration: 1:25:38
Speaker: : Bard Ermentrout
Course:
In this lesson you will learn about fundamental neural phenomena such as oscillations and bursting, and the effects these have on cortical networks.
Difficulty level: Intermediate
Duration: 1:24:30
Speaker: : Bard Ermentrout
Course:
This lesson continues discussing properties of neural oscillations and networks.
Difficulty level: Intermediate
Duration: 1:31:57
Speaker: : Bard Ermentrout
Topics
- Artificial Intelligence (7)
- Philosophy of Science (5)
- Provenance (3)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (8)
- Assembly 2021 (29)
- Brain-hardware interfaces (14)
- Clinical neuroscience (40)
- International Brain Initiative (2)
- (-) Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(62)
- (-) Neuroscience (11)
- Cognitive Science (7)
- (-) Cell signaling (6)
- Brain networks (11)
- (-) Glia (1)
- (-) Electrophysiology (41)
- Learning and memory (5)
- Neuroanatomy (24)
- Neurobiology (16)
- Neurodegeneration (1)
- Neuroimmunology (1)
- Neural networks (15)
- Neurophysiology (27)
- Neuropharmacology (2)
- Neuronal plasticity (16)
- Synaptic plasticity (4)
- Visual system (12)
- Phenome (1)
- General neuroinformatics
(27)
- Computational neuroscience (279)
- Statistics (7)
- Computer Science (21)
- Genomics (34)
- (-)
Data science
(34)
- Open science (61)
- Project management (8)
- Education (4)
- Publishing (4)
- Neuroethics (42)