Displaying 938 results
In this lesson, you will hear about some of the open issues in the field of neuroscience, as well as a discussion about whether neuroscience works, and how can we know?
Difficulty level: Intermediate
Duration: 6:54
Speaker: : Marcus Ghosh
This lesson discusses a gripping neuroscientific question: why have neurons developed the discrete action potential, or spike, as a principle method of communication?
Difficulty level: Intermediate
Duration: 9:34
Speaker: : Dan Goodman
Overview of the evolution of scientific paradigms and the growing role of digital infrastructures in modern neuroscience
Difficulty level: Beginner
Duration: 17:16
Speaker: : Jeanette Hellgren Kotaleski
Mathew Abrams, Director of Science & Training at INCF, outlined how INCF and its global community advance standards, software, infrastructure, and education to enable FAIR and open neuroscience.
Difficulty level: Beginner
Duration: 33:20
Speaker: : Mathew Abrams
EBRAINS is an open research infrastructure which gathering data, tools and computing facilities for brain-related research. In this lecture, professor Trygve Leergaard will introduce the EBRAINS ecosystem and give overview of services for sharing, finding, and using brain research data. He will provide practical examples of how EBRAINS can add values to neuroscience research projects and explain how researchers can engage and utilize the different resources available
Difficulty level: Beginner
Duration: 41:51
Speaker: : Trygve Leergard
This session introduced EBRAINS data and knowledge services and explained how to achieve impactful data sharing through proper curation, FAIR principles, and secure handling of both non-sensitive and sensitive data.
Difficulty level: Beginner
Duration: 25:48
This lesson focused on the importance of organizing data and metadata effectively to ensure that datasets are FAIR and to make data sharing more impactful.
Difficulty level: Beginner
Duration: 14:32
The lesson introduces the Brain Imaging Data Structure (BIDS), the community standard for organizing, curating, and sharing neuroimaging and associated data. The session focuses on understanding the BIDS framework, learning its data structure and validation processes.
Difficulty level: Intermediate
Duration: 38:52
Speaker: : Cyril Pernet
This session moves from BIDS basics into analysis workflows, focusing on how to turn raw, BIDS-organized data into derivatives using BIDS Apps and containers for reproducible processing. It compares end-to-end pipelines across fMRI and PET (and notes EEG/MEG), explains typical preprocessing choices, and shows how standardized inputs plus containerized tools (Docker/AppTainer) yield consistent, auditable outputs.
Difficulty level: Intermediate
Duration: 56:03
Speaker: : Martin Nørgaard
The session explains GDPR rules around data sharing for research in Europe, the distinction between law and ethics, and introduces practical solutions for securely sharing sensitive datasets. Researchers have more flexibility than commonly assumed: scientific research is considered a public interest task, so explicit consent for data sharing isn’t legally required, though transparency and informing participants remain ethically important. The talk also introduces publicneuro.eu, a controlled-access platform that enables sharing neuroimaging datasets with open metadata, DOIs, and customizable access restrictions while ensuring GDPR compliance.
Difficulty level: Intermediate
Duration: 31:12
Speaker: : Cyril Pernet
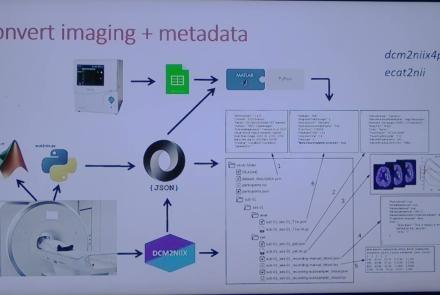
This session introduces the PET-to-BIDS (PET2BIDS) library, a toolkit designed to simplify the conversion and preparation of PET imaging datasets into BIDS-compliant formats. It supports multiple data types and formats (e.g., DICOM, ECAT7+, nifti, JSON), integrates seamlessly with Excel-based metadata, and provides automated routines for metadata updates, blood data conversion, and JSON synchronization. PET2BIDS improves human readability by mapping complex reconstruction names into standardized, descriptive labels and offers extensive documentation, examples, and video tutorials to make adoption easier for researchers.
Difficulty level: Intermediate
Duration: 9:23
Speaker: : Cyril Pernet
This session introduces the PET-to-BIDS (PET2BIDS) library, a toolkit designed to simplify the conversion and preparation of PET imaging datasets into BIDS-compliant formats. It supports multiple data types and formats (e.g., DICOM, ECAT7+, nifti, JSON), integrates seamlessly with Excel-based metadata, and provides automated routines for metadata updates, blood data conversion, and JSON synchronization. PET2BIDS improves human readability by mapping complex reconstruction names into standardized, descriptive labels and offers extensive documentation, examples, and video tutorials to make adoption easier for researchers.
Difficulty level: Intermediate
Duration: 41:04
Speaker: : Martin Nørgaard
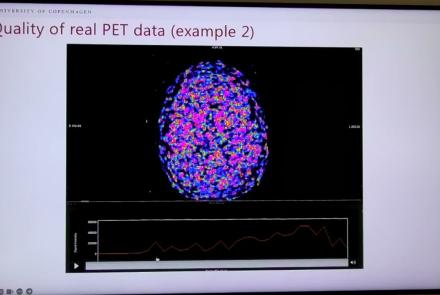
This session dives into practical PET tooling on BIDS data—showing how to run motion correction, register PET↔MRI, extract time–activity curves, and generate standardized PET-BIDS derivatives with clear QC reports. It introduces modular BIDS Apps (head-motion correction, TAC extraction), a full pipeline (PETPrep), and a PET/MRI defacer, with guidance on parameters, outputs, provenance, and why Dockerized containers are the reliable way to run them at scale.
Difficulty level: Intermediate
Duration: 1:05:38
Speaker: : Martin Nørgaard
This session introduces two PET quantification tools—bloodstream for processing arterial blood data and kinfitr for kinetic modeling and quantification—built to work with BIDS/BIDS-derivatives and containers. Bloodstream fuses autosampler and manual measurements (whole blood, plasma, parent fraction) using interpolation or fitted models (incl. hierarchical GAMs) to produce a clean arterial input function (AIF) and whole-blood curves with rich QC reports ready. TAC data (e.g., from PETPrep) and blood (e.g., from bloodstream) can be ingested using kinfitr to run reproducible, GUI-driven analyses: define combined ROIs, calculate weighting factors, estimate blood–tissue delay, choose and chain models (e.g., 2TCM → 1TCM with parameter inheritance), and export parameters/diagnostics. Both are available as Docker apps; workflows emphasize configuration files, reports, and standard outputs to support transparency and reuse.
Difficulty level: Intermediate
Duration: 1:20:56
Speaker: : Granville Matheson
Course:
This lecture gives an introduction to the INCF Short Course: Introduction to Neuroinformatics.
Difficulty level: Beginner
Duration: 34:27
Speaker: : Marja-Leena Linne
Course:
This lecture covers an introduction to connectomics, as well as image processing tools for the study of connectomics.
Difficulty level: Beginner
Duration: 1:23:03
Speaker: : Ignacio Arganda-Carreras
Course:
This lecture covers modeling the neuron in silicon, modeling vision and audition, and sensory fusion using a deep network.
Difficulty level: Beginner
Duration: 1:32:17
Speaker: : Shih-Chii Liu
Course:
This lecture gives an introduction to simulation, models, and the neural simulation tool NEST.
Difficulty level: Beginner
Duration: 1:48:18
Speaker: : Marc-Oliver Gewaltig
Course:
This lecture covers an Introduction to neuron anatomy and signaling, and different types of models, including the Hodgkin-Huxley model.
Difficulty level: Beginner
Duration: 1:23:01
Speaker: : Gaute Einevoll
Course:
This lecture covers the description and characterization of an input-output relationship in a information-theoretic context.
Difficulty level: Beginner
Duration: 1:35:33
Speaker: : Jonathan D. Victor