Lesson type
Difficulty level
This lesson continues with the second workshop on reproducible science, focusing on additional open source tools for researchers and data scientists, such as the R programming language for data science, as well as associated tools like RStudio and R Markdown. Additionally, users are introduced to Python and iPython notebooks, Google Colab, and are given hands-on tutorials on how to create a Binder environment, as well as various containers in Docker and Singularity.
Difficulty level: Beginner
Duration: 1:16:04
Speaker: : Erin Dickie and Sejal Patel
This lecture covers a lot of post-war developments in the science of the mind, focusing first on the cognitive revolution, and concluding with living machines.
Difficulty level: Beginner
Duration: 2:24:35
Speaker: : Paul F.M.J. Verschure
This brief talk goes into work being done at The Alan Turing Institute to solve real-world challenges and democratize computer vision methods to support interdisciplinary and international researchers.
Difficulty level: Beginner
Duration: 7:10
Speaker: : Alden Connor & Beatriz Costa Gomes
This lesson aims to define computational neuroscience in general terms, while providing specific examples of highly successful computational neuroscience projects.
Difficulty level: Beginner
Duration: 59:21
Speaker: : Alla Borisyuk
Course:
This video will document how to run a correlation analysis between the gray matter volume of two different structures using the output from brainlife app-freesurfer-stats.
Difficulty level: Beginner
Duration: 1:33
Speaker: :
Course:
This lecture gives an introduction to simulation, models, and the neural simulation tool NEST.
Difficulty level: Beginner
Duration: 1:48:18
Speaker: : Marc-Oliver Gewaltig
Course:
This lecture covers an Introduction to neuron anatomy and signaling, and different types of models, including the Hodgkin-Huxley model.
Difficulty level: Beginner
Duration: 1:23:01
Speaker: : Gaute Einevoll
This lecture covers an Introduction to neuron anatomy and signaling, and different types of models, including the Hodgkin-Huxley model.
Difficulty level: Beginner
Duration: 1:23:01
Speaker: : Gaute Einevoll
This lesson discuses forms of neural plasticity on many levels, including short-term, long-term, metaplasticity, and structural plasticity. During the lesson you will also be presented with examples related to the modelling of biochemical networks.
Difficulty level: Beginner
Duration: 1:11:29
Speaker: : Upi Bhalla
This lesson provides an introduction to modelling of chemical computation in the brain.
Difficulty level: Beginner
Duration: 1:00:11
Speaker: : Upi Bhalla
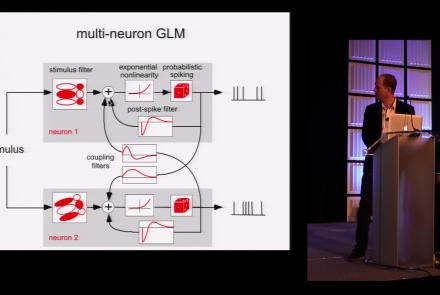
This lesson is part 1 of 2 of a tutorial on statistical models for neural data.
Difficulty level: Beginner
Duration: 1:45:48
Speaker: : Jonathan Pillow
This lesson is part 2 of 2 of a tutorial on statistical models for neural data.
Difficulty level: Beginner
Duration: 1:50:31
Speaker: : Jonathan Pillow
Course:
This lecture covers an Introduction to neuron anatomy and signaling, as well as different types of models, including the Hodgkin-Huxley model.
Difficulty level: Beginner
Duration: 1:23:01
Speaker: : Gaute Einevoll
Course:
This lecture describes forms of plasticity on many levels: short-term, long-term, metaplasticity, and structural plasticity. Included in this lecture are also examples related to modelling of biochemical networks.
Difficulty level: Beginner
Duration: 1:11:29
Speaker: : Upi Bhalla
Course:
This lesson provides an introduction to modelling of chemical computation in the brain.
Difficulty level: Beginner
Duration: 1:00:11
Speaker: : Upi Bhalla
This lesson provides an introduction to the role of models in theoretical neuroscience, particularly focusing on David Marr's work on levels of description/analysis of the brain as a complex system: computation, algorithm & representation, and implementation.
Difficulty level: Beginner
Duration: 19:26
Speaker: : Jakob Macke
In this lesson, you will learn about different types of models, model complexity, and how to choose an appropriate model.
Difficulty level: Beginner
Duration: 39:09
Speaker: : Astrid Prinz
This lesson provides an overview of balanced excitatory-inhibitory (E-I) networks, stability, and gain modulation.
Difficulty level: Beginner
Duration: 1:22:11
Speaker: : Kenneth Miller
This lesson introduces methods for dimensionality reduction of data, with focus on factor analysis.
Difficulty level: Beginner
Duration: 1:16:47
Speaker: : Byron Yu
This lecture delves into the dynamics of neural computation, from the spiking activity of single neurons to regional cortical population coding and network activity.
Difficulty level: Beginner
Duration: 1:39:32
Speaker: : Julijana Gjorgjieva
Topics
- Artificial Intelligence (6)
- Philosophy of Science (5)
- Provenance (2)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (6)
- Assembly 2021 (29)
- Brain-hardware interfaces (13)
- Clinical neuroscience (17)
- International Brain Initiative (2)
- Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(45)
- Neuroscience (9)
- Cognitive Science (7)
- Cell signaling (3)
- Brain networks (4)
- Glia (1)
- Electrophysiology (16)
- Learning and memory (3)
- Neuroanatomy (17)
- Neurobiology (7)
- Neurodegeneration (1)
- Neuroimmunology (1)
- Neural networks (4)
- Neurophysiology (22)
- Neuropharmacology (2)
- Synaptic plasticity (2)
- Visual system (12)
- Phenome (1)
- General neuroinformatics
(15)
- (-) Computational neuroscience (192)
- (-) Statistics (2)
- Computer Science (15)
- Genomics (25)
- Data science
(24)
- Open science (52)
- Project management (7)
- Education (3)
- Publishing (4)
- Neuroethics (35)