Manipulate the default connectome provided with TVB to see how structural lesions effect brain dynamics. In this hands-on session you will insert lesions into the connectome within the TVB graphical user interface (GUI). Afterwards, the modified connectome will be used for simulations and the resulting activity will be analysed using functional connectivity.
Difficulty level: Beginner
Duration: 31:22
Speaker: : Paul Triebkorn
This presentation discusses the impact of data sharing in stroke.
Difficulty level: Intermediate
Duration: 16:33
Speaker: : Valeria Caso
This talks presents an overview of the potential for data federation in stroke research.
Difficulty level: Intermediate
Duration: 21:37
Speaker: : Maurizio A. Leone
Course:
This talk focuses on the EAN Scientific Panel of Stroke, in particular on the aims and roles of the panel.
Difficulty level: Intermediate
Duration: 18:19
Speaker: : Anna Bersano
This lecture provides an introduction to optogenetics, a biological technique to control the activity of neurons or other cell types with light.
Difficulty level: Beginner
Duration: 39:34
Speaker: : Adam Packer
Course:
This primer on optogenetics primer discusses how to manipulate neuronal populations with light at millisecond resolution and offers possible applications such as curing the blind and "playing the piano" with cortical neurons.
Difficulty level: Beginner
Duration: 59:06
Speaker: : Clay Reid
Learn how to create a standard extracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 23:10
Speaker: : Ryan Ly
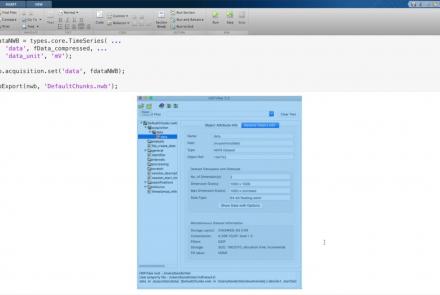
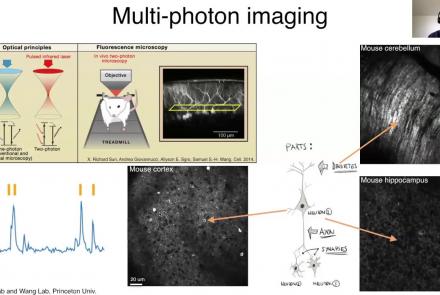
Learn how to create a standard calcium imaging dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 31:04
Speaker: : Ryan Ly
In this tutorial, you will learn how to create a standard intracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 20:23
Speaker: : Pamela Baker
In this tutorial, you will learn how to use the icephys-metadata extension to enter meta-data detailing your experimental paradigm.
Difficulty level: Intermediate
Duration: 27:18
Speaker: : Oliver Ruebel
This lesson provides instructions on how to build and share extensions in NWB.
Difficulty level: Advanced
Duration: 20:29
Speaker: : Ryan Ly
Learn how to build custom APIs for extension.
Difficulty level: Advanced
Duration: 25:40
Speaker: : Andrew Tritt
This lesson provides instruction on advanced writing strategies in HDF5 that are accessible through PyNWB.
Difficulty level: Advanced
Duration: 23:00
Speaker: : Oliver Ruebel
In this tutorial, users learn how to create a standard extracellular electrophysiology dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 45:46
Speaker: : Ben Dichter
Learn how to create a standard calcium imaging dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 39:10
Speaker: : Ben Dichter
Learn how to create a standard intracellular electrophysiology dataset in NWB.
Difficulty level: Intermediate
Duration: 20:22
Speaker: : Pamela Baker
This lesson provides a tutorial on how to handle writing very large data in MatNWB.
Difficulty level: Advanced
Duration: 16:18
Speaker: : Ben Dichter
This lesson gives an overview of the Brainstorm package for analyzing extracellular electrophysiology, including preprocessing, spike sorting, trial alignment, and spectrotemporal decomposition.
Difficulty level: Intermediate
Duration: 47:47
Speaker: : Konstantinos Nasiotis
This lesson provides an overview of the CaImAn package, as well as a demonstration of usage with NWB.
Difficulty level: Intermediate
Duration: 44:37
Speaker: : Andrea Giovannucci
This lesson gives an overview of the SpikeInterface package, including demonstration of data loading, preprocessing, spike sorting, and comparison of spike sorters.
Difficulty level: Intermediate
Duration: 1:10:28
Speaker: : Alessio Buccino
Topics
- Artificial Intelligence (7)
- Philosophy of Science (5)
- Provenance (3)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (8)
- Assembly 2021 (29)
- Brain-hardware interfaces (14)
- Clinical neuroscience (40)
- International Brain Initiative (2)
- Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(62)
- Neuroscience (11)
- Cognitive Science (7)
- (-) Cell signaling (6)
- Brain networks (11)
- (-) Glia (1)
- Electrophysiology (41)
- Learning and memory (5)
- (-) Neuroanatomy (24)
- Neurobiology (16)
- Neurodegeneration (1)
- Neuroimmunology (1)
- Neural networks (15)
- Neurophysiology (27)
- Neuropharmacology (2)
- Neuronal plasticity (16)
- Synaptic plasticity (4)
- Visual system (12)
- Phenome (1)
- General neuroinformatics
(27)
- Computational neuroscience (276)
- Statistics (7)
- Computer Science (20)
- Genomics (33)
- Data science
(33)
- Open science (57)
- Project management (8)
- Education (4)
- Publishing (4)
- Neuroethics (40)