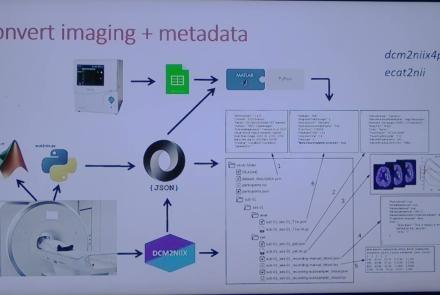
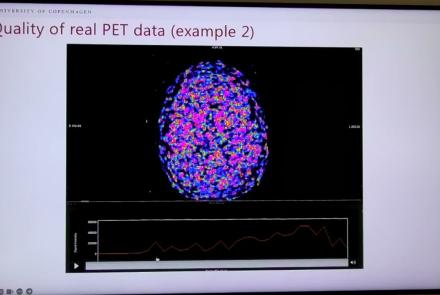
This lightning talk describes an automated pipline for positron emission tomography (PET) data.
Difficulty level: Intermediate
Duration: 7:27
Speaker: : Soodeh Moallemian
This session introduces the PET-to-BIDS (PET2BIDS) library, a toolkit designed to simplify the conversion and preparation of PET imaging datasets into BIDS-compliant formats. It supports multiple data types and formats (e.g., DICOM, ECAT7+, nifti, JSON), integrates seamlessly with Excel-based metadata, and provides automated routines for metadata updates, blood data conversion, and JSON synchronization. PET2BIDS improves human readability by mapping complex reconstruction names into standardized, descriptive labels and offers extensive documentation, examples, and video tutorials to make adoption easier for researchers.
Difficulty level: Intermediate
Duration: 9:23
Speaker: : Cyril Pernet
This session introduces the PET-to-BIDS (PET2BIDS) library, a toolkit designed to simplify the conversion and preparation of PET imaging datasets into BIDS-compliant formats. It supports multiple data types and formats (e.g., DICOM, ECAT7+, nifti, JSON), integrates seamlessly with Excel-based metadata, and provides automated routines for metadata updates, blood data conversion, and JSON synchronization. PET2BIDS improves human readability by mapping complex reconstruction names into standardized, descriptive labels and offers extensive documentation, examples, and video tutorials to make adoption easier for researchers.
Difficulty level: Intermediate
Duration: 41:04
Speaker: : Martin Nørgaard
This session dives into practical PET tooling on BIDS data—showing how to run motion correction, register PET↔MRI, extract time–activity curves, and generate standardized PET-BIDS derivatives with clear QC reports. It introduces modular BIDS Apps (head-motion correction, TAC extraction), a full pipeline (PETPrep), and a PET/MRI defacer, with guidance on parameters, outputs, provenance, and why Dockerized containers are the reliable way to run them at scale.
Difficulty level: Intermediate
Duration: 1:05:38
Speaker: : Martin Nørgaard
This session introduces two PET quantification tools—bloodstream for processing arterial blood data and kinfitr for kinetic modeling and quantification—built to work with BIDS/BIDS-derivatives and containers. Bloodstream fuses autosampler and manual measurements (whole blood, plasma, parent fraction) using interpolation or fitted models (incl. hierarchical GAMs) to produce a clean arterial input function (AIF) and whole-blood curves with rich QC reports ready. TAC data (e.g., from PETPrep) and blood (e.g., from bloodstream) can be ingested using kinfitr to run reproducible, GUI-driven analyses: define combined ROIs, calculate weighting factors, estimate blood–tissue delay, choose and chain models (e.g., 2TCM → 1TCM with parameter inheritance), and export parameters/diagnostics. Both are available as Docker apps; workflows emphasize configuration files, reports, and standard outputs to support transparency and reuse.
Difficulty level: Intermediate
Duration: 1:20:56
Speaker: : Granville Matheson
This lecture gives an introduction to the European Academy of Neurology, its recent achievements and ambitions.
Difficulty level: Intermediate
Duration: 21:57
Speaker: : Paul Boon
This lecture discusses the the importance and need for data sharing in clinical neuroscience.
Difficulty level: Intermediate
Duration: 25:22
Speaker: : Thomas Berger
This lecture presents the Medical Informatic Platform's data federation for Traumatic Brain Injury.
Difficulty level: Intermediate
Duration: 25:55
Speaker: : Stefano Finazzi
This lecture gives an overview on the European Health Dataspace.
Difficulty level: Intermediate
Duration: 26:33
Speaker: : Licino Kustra Mano
This lecture presents the Medical Informatics Platform's data federation in epilepsy.
Difficulty level: Intermediate
Duration: 27:09
Speaker: : Philippe Ryvlin
This lesson describes how DataLad allows you to track and mange both your data and analysis code, thereby facilitating reliable, reproducible, and shareable research.
Difficulty level: Intermediate
Duration: 59:34
Speaker: : Yaroslav O. Halchenko
Course:
The International Brain Initiative (IBI) is a consortium of the world’s major large-scale brain initiatives and other organizations with a vested interest in catalyzing and advancing neuroscience research through international collaboration and knowledge sharing. This workshop introduces the IBI, the efforts of the Data Standards and Sharing Working Group, and keynote lectures on the impact of data standards and sharing on large-scale brain projects, as well as a discussion on prospects and needs for neural data sharing.
Difficulty level: Intermediate
Duration: 2:06:58
This lecture covers how you can make your data public through EBRAINS. This talk focuses on the ethical considerations for sharing data, the requirements that are imposed by various regulations, particularly for sharing human data. The lecture also includes a discussion of how EBRAINS designs its services to deal with the ethical and regulatory aspects of sharing these kinds of data.
Difficulty level: Intermediate
Duration: 16:15
Speaker: : Maaike van Swieten and Jan Bjaalie
This lecture discusses differential privacy and synthetic data in the context of medical data sharing in clinical neurosciences.
Difficulty level: Intermediate
Duration: 20:26
Speaker: : Minos Garofalakis
This presentation discusses the impact of data sharing in stroke.
Difficulty level: Intermediate
Duration: 16:33
Speaker: : Valeria Caso
This talks discusses data sharing in the context of dementia. It explains why data sharing in dementia is important, how data is usually shared in the field and illustrates two examples: the Netherlands Consortium Dementia cohorts and the European Platform for Neurodegenerative Diseases.
Difficulty level: Intermediate
Duration: 21:21
Speaker: : Pieter Jelle Visser
This talk introduces data sharing initiatives in Epilepsy, particularly across Europe.
Difficulty level: Intermediate
Duration: 13:56
Speaker: : J. Helen Cross
Topics
- Artificial Intelligence (1)
- Notebooks (1)
- Provenance (1)
- DANDI archive (1)
- EBRAINS RI (6)
- Animal models (2)
- Brain-hardware interfaces (1)
- Clinical neuroscience (23)
- General neuroscience
(17)
- General neuroinformatics
(1)
- Computational neuroscience (81)
- Statistics (5)
- Computer Science (5)
- Genomics (8)
- Data science
(10)
- Open science (5)
- Project management (1)
- Education (1)
- Neuroethics (5)