In this lightning talk, you will learn about BrainGlobe, an initiative which exists to facilitate the development of interoperable Python-based tools for computational neuroanatomy.
Difficulty level: Beginner
Duration: 3:33
Speaker: : Alessandro Felder
This lesson consists of a panel discussion, wrapping up the INCF Neuroinformatics Assembly 2023 workshop Research Workflows for Collaborative Neuroscience.
Difficulty level: Beginner
Duration: 25:33
Speaker: :
This talk gives an overview of the complicated nature of sharing of neuroscientific data in an environment of numerous and often conflicting legal systems around the world.
Difficulty level: Beginner
Duration: 14:00
Speaker: : Franco Pestilli
This talk provides an overview of the FAIR-aligned efforts of MATLAB and MathWorks, from the technological building blocks to the open science coordination involved in facilitating greater transparency and efficiency in neuroscience and neuroinformatics.
Difficulty level: Beginner
Duration: 15:41
Speaker: : Vijay Iyer
This lesson aims to define computational neuroscience in general terms, while providing specific examples of highly successful computational neuroscience projects.
Difficulty level: Beginner
Duration: 59:21
Speaker: : Alla Borisyuk
Course:
This brief video provides an introduction to brainlife.io, a free cloud computing platform for neuroimaging data analysis.
Difficulty level: Beginner
Duration: 2:41
Speaker: :
Course:
This video will document the process of uploading data into a brainlife project using ezBIDS.
Difficulty level: Beginner
Duration: 6:15
Speaker: :
Course:
This lesson visually documents the process of uploading data to brainlife via the command line interface (CLI).
Difficulty level: Beginner
Duration: 1:28
Speaker: :
Course:
This video will document the process of visualizing the provenance of each step performed to generate a data object on brainlife.
Difficulty level: Beginner
Duration: 0:21
Speaker: :
Course:
This video will document the process of downloading and running the "reproduce.sh" script, which will automatically run all of the steps to generate a data object locally on a user's machine.
Difficulty level: Beginner
Duration: 3:44
Speaker: :
Course:
This brief video walks you through the steps necessary when creating a project on brainlife.io.
Difficulty level: Beginner
Duration: 1:45
Speaker: :
Course:
This brief video rus through how to make an accout on brainlife.io.
Difficulty level: Beginner
Duration: 0:30
Speaker: :
Course:
This short video shows how data in a brainlife.io publication can be opened from a DOI inside a published article. The video provides an example of how the DOI deposited on the journal can be opened with a web browser to redirect to the associated data publication on brainlife.io.
Difficulty level: Beginner
Duration: 2:18
Speaker: :
Course:
This video will document the process of importing a dataset archived on OpenNeuro from the Datasets tab into a brainlife project.
Difficulty level: Beginner
Duration: 1:06
Speaker: :
Course:
This lecture gives an introduction to simulation, models, and the neural simulation tool NEST.
Difficulty level: Beginner
Duration: 1:48:18
Speaker: : Marc-Oliver Gewaltig
Course:
This lecture covers an Introduction to neuron anatomy and signaling, and different types of models, including the Hodgkin-Huxley model.
Difficulty level: Beginner
Duration: 1:23:01
Speaker: : Gaute Einevoll
Course:
This lecture covers structured data, databases, federating neuroscience-relevant databases, and ontologies.
Difficulty level: Beginner
Duration: 1:30:45
Speaker: : Maryann Martone
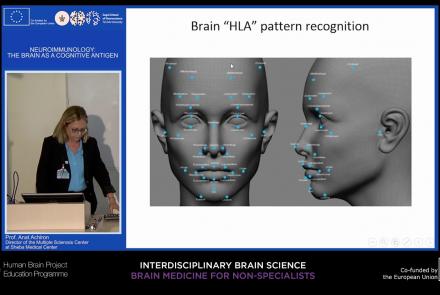
This lecture focuses on how the immune system can target and attack the nervous system to produce autoimmune responses that may result in diseases such as multiple sclerosis, neuromyelitis, and lupus cerebritis manifested by motor, sensory, and cognitive impairments. Despite the fact that the brain is an immune-privileged site, autoreactive lymphocytes producing proinflammatory cytokines can cause active brain inflammation, leading to myelin and axonal loss.
Difficulty level: Beginner
Duration: 37:36
Speaker: : Anat Achiron
This lesson discusses both state-of-the-art detection and prevention schema in working with neurodegenerative diseases.
Difficulty level: Beginner
Duration: 1:02:29
Speaker: : Nir Giladi
This lecture consists of the second half of the introduction to signal transduction, here focusing on cell receptors and signalling cascades.
Difficulty level: Beginner
Duration: 41:38
Speaker: : Christoph Schwarzer
Topics
- (-) Artificial Intelligence (6)
- Philosophy of Science (5)
- Provenance (2)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (6)
- Assembly 2021 (29)
- Brain-hardware interfaces (13)
- Clinical neuroscience (17)
- International Brain Initiative (2)
- Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(45)
- Neuroscience (9)
- Cognitive Science (7)
- (-) Cell signaling (3)
- Brain networks (4)
- Glia (1)
- Electrophysiology (16)
- Learning and memory (3)
- Neuroanatomy (17)
- Neurobiology (7)
- (-) Neurodegeneration (1)
- (-) Neuroimmunology (1)
- Neural networks (4)
- Neurophysiology (22)
- Neuropharmacology (2)
- (-) Synaptic plasticity (2)
- (-) Visual system (12)
- Phenome (1)
- General neuroinformatics
(15)
- (-) Computational neuroscience (195)
- Statistics (2)
- Computer Science (15)
- Genomics (26)
- Data science
(24)
- (-) Open science (56)
- Project management (7)
- Education (3)
- Publishing (4)
- Neuroethics (37)