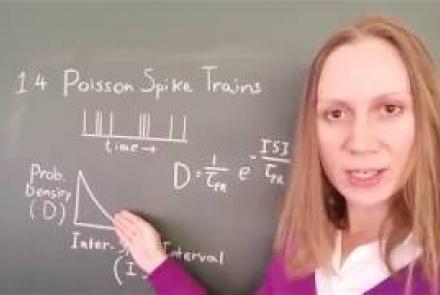Course:
In addition to outlining the motivations behind preprocessing EEG data in general, this lesson covers the first step in preprocessing data with EEGLAB, importing raw data.
Difficulty level: Beginner
Duration: 8:30
Speaker: : Arnaud Delorme
Course:
Continuing along the EEGLAB preprocessing pipeline, this tutorial walks users through how to import data events as well as EEG channel locations.
Difficulty level: Beginner
Duration: 11:53
Speaker: : Arnaud Delorme
Course:
This tutorial demonstrates how to re-reference and resample raw data in EEGLAB, why such steps are important or useful in the preprocessing pipeline, and how choices made at this step may affect subsequent analyses.
Difficulty level: Beginner
Duration: 11:48
Speaker: : Arnaud Delorme
Course:
In this tutorial, users learn about the various filtering options in EEGLAB, how to inspect channel properties for noisy signals, as well as how to filter out specific components of EEG data (e.g., electrical line noise).
Difficulty level: Beginner
Duration: 10:46
Speaker: : Arnaud Delorme
Course:
This tutorial instructs users how to visually inspect partially pre-processed neuroimaging data in EEGLAB, specifically how to use the data browser to investigate specific channels, epochs, or events for removable artifacts, biological (e.g., eye blinks, muscle movements, heartbeat) or otherwise (e.g., corrupt channel, line noise).
Difficulty level: Beginner
Duration: 5:08
Speaker: : Arnaud Delorme
Course:
This tutorial provides instruction on how to use EEGLAB to further preprocess EEG datasets by identifying and discarding bad channels which, if left unaddressed, can corrupt and confound subsequent analysis steps.
Difficulty level: Beginner
Duration: 13:01
Speaker: : Arnaud Delorme
Course:
Users following this tutorial will learn how to identify and discard bad EEG data segments using the MATLAB toolbox EEGLAB.
Difficulty level: Beginner
Duration: 11:25
Speaker: : Arnaud Delorme
Course:
An introduction to data management, manipulation, visualization, and analysis for neuroscience. Students will learn scientific programming in Python, and use this to work with example data from areas such as cognitive-behavioral research, single-cell recording, EEG, and structural and functional MRI. Basic signal processing techniques including filtering are covered. The course includes a Jupyter Notebook and video tutorials.
Difficulty level: Beginner
Duration: 1:09:16
Speaker: : Aaron J. Newman
This lesson describes spike timing-dependent plasticity (STDP), a biological process that adjusts the strength of connections between neurons in the brain, and how one can implement or mimic this process in a computational model. You will also find links for practical exercises at the bottom of this page.
Difficulty level: Intermediate
Duration: 12:50
Speaker: : Dan Goodman
This lesson provides a brief introduction to the Computational Modeling of Neuronal Plasticity.
Difficulty level: Intermediate
Duration: 0:40
Speaker: : Florence I. Kleberg
In this lesson, you will be introducted to a type of neuronal model known as the leaky integrate-and-fire (LIF) model.
Difficulty level: Intermediate
Duration: 1:23
Speaker: : Florence I. Kleberg
This lesson goes over various potential inputs to neuronal synapses, loci of neural communication.
Difficulty level: Intermediate
Duration: 1:20
Speaker: : Florence I. Kleberg
This lesson describes the how and why behind implementing integration time steps as part of a neuronal model.
Difficulty level: Intermediate
Duration: 1:08
Speaker: : Florence I. Kleberg
In this lesson, you will learn about neural spike trains which can be characterized as having a Poisson distribution.
Difficulty level: Intermediate
Duration: 1:18
Speaker: : Florence I. Kleberg
This lesson covers spike-rate adaptation, the process by which a neuron's firing pattern decays to a low, steady-state frequency during the sustained encoding of a stimulus.
Difficulty level: Intermediate
Duration: 1:26
Speaker: : Florence I. Kleberg
This lesson provides a brief explanation of how to implement a neuron's refractory period in a computational model.
Difficulty level: Intermediate
Duration: 0:42
Speaker: : Florence I. Kleberg
In this lesson, you will learn a computational description of the process which tunes neuronal connectivity strength, spike-timing-dependent plasticity (STDP).
Difficulty level: Intermediate
Duration: 2:40
Speaker: : Florence I. Kleberg
This lesson reviews theoretical and mathematical descriptions of correlated spike trains.
Difficulty level: Intermediate
Duration: 2:54
Speaker: : Florence I. Kleberg
This lesson investigates the effect of correlated spike trains on spike-timing dependent plasticity (STDP).
Difficulty level: Intermediate
Duration: 1:43
Speaker: : Florence I. Kleberg
This lesson goes over synaptic normalisation, the homeostatic process by which groups of weighted inputs scale up or down their biases.
Difficulty level: Intermediate
Duration: 2:58
Speaker: : Florence I. Kleberg
Topics
- Artificial Intelligence (7)
- Philosophy of Science (5)
- Provenance (3)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (8)
- Assembly 2021 (29)
- Brain-hardware interfaces (14)
- Clinical neuroscience (40)
- International Brain Initiative (2)
- Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(62)
- (-) Neuroscience (11)
- Cognitive Science (7)
- Cell signaling (6)
- Brain networks (11)
- (-) Glia (1)
- Electrophysiology (41)
- Learning and memory (5)
- Neuroanatomy (24)
- (-) Neurobiology (16)
- Neurodegeneration (1)
- Neuroimmunology (1)
- Neural networks (15)
- Neurophysiology (27)
- Neuropharmacology (2)
- (-) Neuronal plasticity (16)
- Synaptic plasticity (4)
- Visual system (12)
- Phenome (1)
- General neuroinformatics
(27)
- Computational neuroscience (275)
- Statistics (7)
- Computer Science (20)
- Genomics (32)
- Data science
(33)
- Open science (57)
- (-) Project management (8)
- Education (4)
- Publishing (4)
- (-) Neuroethics (40)