Manipulate the default connectome provided with TVB to see how structural lesions effect brain dynamics. In this hands-on session you will insert lesions into the connectome within the TVB graphical user interface (GUI). Afterwards, the modified connectome will be used for simulations and the resulting activity will be analysed using functional connectivity.
Difficulty level: Beginner
Duration: 31:22
Speaker: : Paul Triebkorn
This presentation discusses the impact of data sharing in stroke.
Difficulty level: Intermediate
Duration: 16:33
Speaker: : Valeria Caso
This talks presents an overview of the potential for data federation in stroke research.
Difficulty level: Intermediate
Duration: 21:37
Speaker: : Maurizio A. Leone
Course:
This talk focuses on the EAN Scientific Panel of Stroke, in particular on the aims and roles of the panel.
Difficulty level: Intermediate
Duration: 18:19
Speaker: : Anna Bersano
Course:
The state of the field regarding the diagnosis and treatment of major depressive disorder (MDD) is discussed. Current challenges and opportunities facing the research and clinical communities are outlined, including appropriate quantitative and qualitative analyses of the heterogeneity of biological, social, and psychiatric factors which may contribute to MDD.
Difficulty level: Beginner
Duration: 1:29:28
Speaker: : Brett Jones, Victor Tang
This lesson gives a description of the BrainHealth Databank, a repository of many types of health-related data, whose aim is to accelerate research, improve care, and to help better understand and diagnose mental illness, as well as develop new treatments and prevention strategies.
This lesson corresponds to slides 46-78 of the PDF below.
Difficulty level: Beginner
Duration: 1:12:25
Speaker: : Joanna Yu
This lesson describes not only the need for precision medicine, but also the current state of the methods, pharmacogenetic approaches, utility and implementation of such care today.
This lesson corresponds to slides 1-50 of the PowerPoint below.
Difficulty level: Beginner
Duration: 1:24:30
Speaker: : Dan Felsky
This lecture discusses what defines an integrative approach regarding research and methods, including various study designs and models which are appropriate choices when attempting to bridge data domains; a necessity when whole-person modelling.
Difficulty level: Beginner
Duration: 1:28:14
Speaker: : Dan Felsky
This is the Introductory Module to the Deep Learning Course at CDS, a course that covered the latest techniques in deep learning and representation learning, focusing on supervised and unsupervised deep learning, embedding methods, metric learning, convolutional and recurrent nets, with applications to computer vision, natural language understanding, and speech recognition.
Difficulty level: Intermediate
Duration: 50:17
Speaker: : Yann LeCun and Alfredo Canziani
This module covers the concepts of gradient descent and the backpropagation algorithm and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:51:03
Speaker: : Yann LeCun
This lecture covers concepts associated with neural nets, including rotation and squashing, and is a part of the Deep Learning Course at New York University's Center for Data Science (CDS).
Difficulty level: Intermediate
Duration: 1:01:53
Speaker: : Alfredo Canziani
This lesson provides a detailed description of some of the modules and architectures involved in the development of neural networks.
Difficulty level: Intermediate
Duration: 1:42:26
Speaker: : Yann LeCun and Alfredo Canziani
This lecture covers the concept of neural nets training (tools, classification with neural nets, and PyTorch implementation) and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:05:47
Speaker: : Alfredo Canziani
This lecture covers the concept of parameter sharing: recurrent and convolutional nets and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:59:47
Speaker: : Yann LeCun and Alfredo Canziani
This lecture covers the concept of convolutional nets in practice and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 51:40
Speaker: : Yann LeCun
This lecture discusses the concept of natural signals properties and the convolutional nets in practice and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:09:12
Speaker: : Alfredo Canziani
This lecture covers the concept of recurrent neural networks: vanilla and gated (LSTM) and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:05:36
Speaker: : Alfredo Canziani
This lecture is a foundationational lecture for the concept of energy-based models with a particular focus on the joint embedding method and latent variable energy-based models (LV-EBMs) and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:51:30
Speaker: : Yann LeCun
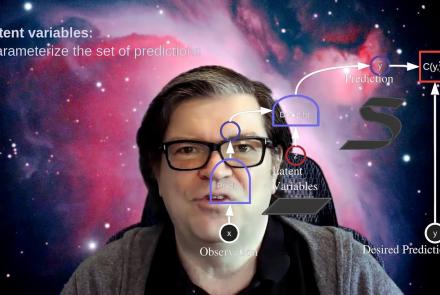
This lecture covers the concept of inference in latent variable energy based models (LV-EBMs) and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:01:04
Speaker: : Alfredo Canziani
This panel discussion covers how energy based models are used and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 10:42
Speaker: : Yann LeCun and Alfredo Canziani
Topics
- Artificial Intelligence (7)
- Philosophy of Science (5)
- Provenance (3)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (8)
- Assembly 2021 (29)
- Brain-hardware interfaces (14)
- Clinical neuroscience (40)
- International Brain Initiative (2)
- Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(62)
- Neuroscience (11)
- Cognitive Science (7)
- Cell signaling (6)
- Brain networks (11)
- (-) Glia (1)
- Electrophysiology (41)
- Learning and memory (5)
- (-) Neuroanatomy (24)
- (-) Neurobiology (16)
- Neurodegeneration (1)
- (-) Neuroimmunology (1)
- Neural networks (15)
- (-) Neurophysiology (27)
- Neuropharmacology (2)
- Neuronal plasticity (16)
- Synaptic plasticity (4)
- Visual system (12)
- Phenome (1)
- General neuroinformatics
(27)
- Computational neuroscience (279)
- Statistics (7)
- Computer Science (21)
- Genomics (34)
- Data science
(34)
- Open science (61)
- Project management (8)
- Education (4)
- Publishing (4)
- (-) Neuroethics (42)