Lesson type
Difficulty level
This talk describes the NIH-funded SPARC Data Structure, and how this project navigates ontology development while keeping in mind the FAIR science principles.
Difficulty level: Beginner
Duration: 25:44
Speaker: : Fahim Imam
This lesson provides an overview of the current status in the field of neuroscientific ontologies, presenting examples of data organization and standards, particularly from neuroimaging and electrophysiology.
Difficulty level: Intermediate
Duration: 33:41
Speaker: : Yaroslav O. Halchenko
This lesson continues from part one of the lecture Ontologies, Databases, and Standards, diving deeper into a description of ontologies and knowledg graphs.
Difficulty level: Intermediate
Duration: 50:18
Speaker: : Jeff Grethe
Course:
This lecture covers structured data, databases, federating neuroscience-relevant databases, and ontologies.
Difficulty level: Beginner
Duration: 1:30:45
Speaker: : Maryann Martone
Course:
This lecture covers FAIR atlases, including their background and construction, as well as how they can be created in line with the FAIR principles.
Difficulty level: Beginner
Duration: 14:24
Speaker: : Heidi Kleven
This lecture focuses on ontologies for clinical neurosciences.
Difficulty level: Intermediate
Duration: 21:54
Speaker: : Martin Hofmann-Apitius
This lecture provides an introduction to optogenetics, a biological technique to control the activity of neurons or other cell types with light.
Difficulty level: Beginner
Duration: 39:34
Speaker: : Adam Packer
Course:
This primer on optogenetics primer discusses how to manipulate neuronal populations with light at millisecond resolution and offers possible applications such as curing the blind and "playing the piano" with cortical neurons.
Difficulty level: Beginner
Duration: 59:06
Speaker: : Clay Reid
Learn how to create a standard extracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 23:10
Speaker: : Ryan Ly
Learn how to create a standard calcium imaging dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 31:04
Speaker: : Ryan Ly
In this tutorial, you will learn how to create a standard intracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 20:23
Speaker: : Pamela Baker
In this tutorial, you will learn how to use the icephys-metadata extension to enter meta-data detailing your experimental paradigm.
Difficulty level: Intermediate
Duration: 27:18
Speaker: : Oliver Ruebel
This lesson provides instructions on how to build and share extensions in NWB.
Difficulty level: Advanced
Duration: 20:29
Speaker: : Ryan Ly
Learn how to build custom APIs for extension.
Difficulty level: Advanced
Duration: 25:40
Speaker: : Andrew Tritt
This lesson provides instruction on advanced writing strategies in HDF5 that are accessible through PyNWB.
Difficulty level: Advanced
Duration: 23:00
Speaker: : Oliver Ruebel
In this tutorial, users learn how to create a standard extracellular electrophysiology dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 45:46
Speaker: : Ben Dichter
Learn how to create a standard calcium imaging dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 39:10
Speaker: : Ben Dichter
Learn how to create a standard intracellular electrophysiology dataset in NWB.
Difficulty level: Intermediate
Duration: 20:22
Speaker: : Pamela Baker
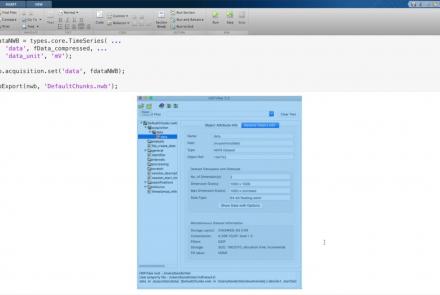
This lesson provides a tutorial on how to handle writing very large data in MatNWB.
Difficulty level: Advanced
Duration: 16:18
Speaker: : Ben Dichter
This lesson gives an overview of the Brainstorm package for analyzing extracellular electrophysiology, including preprocessing, spike sorting, trial alignment, and spectrotemporal decomposition.
Difficulty level: Intermediate
Duration: 47:47
Speaker: : Konstantinos Nasiotis
Topics
- Artificial Intelligence (7)
- Philosophy of Science (5)
- Provenance (3)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (8)
- Assembly 2021 (29)
- Brain-hardware interfaces (14)
- Clinical neuroscience (40)
- International Brain Initiative (2)
- Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(62)
- Neuroscience (11)
- Cognitive Science (7)
- Cell signaling (6)
- Brain networks (11)
- Glia (1)
- Electrophysiology (41)
- Learning and memory (5)
- (-) Neuroanatomy (24)
- Neurobiology (16)
- Neurodegeneration (1)
- Neuroimmunology (1)
- Neural networks (15)
- Neurophysiology (27)
- Neuropharmacology (2)
- Neuronal plasticity (16)
- Synaptic plasticity (4)
- Visual system (12)
- Phenome (1)
- General neuroinformatics
(27)
- Computational neuroscience (275)
- Statistics (7)
- Computer Science (20)
- (-) Genomics (32)
- Data science
(33)
- Open science (57)
- Project management (8)
- Education (4)
- Publishing (4)
- Neuroethics (40)