Course:
In this lesson, while learning about the need for increased large-scale collaborative science that is transparent in nature, users also are given a tutorial on using Synapse for facilitating reusable and reproducible research.
Difficulty level: Beginner
Duration: 1:15:12
Speaker: : Abhi Pratap
This lesson provides an overview of how to conceptualize, design, implement, and maintain neuroscientific pipelines in via the cloud-based computational reproducibility platform Code Ocean.
Difficulty level: Beginner
Duration: 17:01
Speaker: : David Feng
This lesson provides an overview of how to construct computational pipelines for neurophysiological data using DataJoint.
Difficulty level: Beginner
Duration: 17:37
Speaker: : Dimitri Yatsenko
This hands-on tutorial walks you through DataJoint platform, highlighting features and schema which can be used to build robost neuroscientific pipelines.
Difficulty level: Beginner
Duration: 26:06
Speaker: : Milagros Marin
This lesson provides an introduction to the DataLad, a free and open source distributed data management system that keeps track of your data, creates structure, ensures reproducibility, supports collaboration, and integrates with widely used data infrastructure.
Difficulty level: Beginner
Duration: 22:56
Speaker: : Michał Szczepanik
This lesson introduces several open science tools like Docker and Apptainer which can be used to develop portable and reproducible software environments.
Difficulty level: Beginner
Duration: 17:22
Speaker: : Joanes Grandjean
This lecture provides a detailed description of how to incorporate HED annotation into your neuroimaging data pipeline.
Difficulty level: Beginner
Duration: 33:36
Speaker: : Dung Truong
This lecture covers a wide range of aspects regarding neuroinformatics and data governance, describing both their historical developments and current trajectories. Particular tools, platforms, and standards to make your research more FAIR are also discussed.
Difficulty level: Beginner
Duration: 54:58
Speaker: : Franco Pestilli
Manipulate the default connectome provided with TVB to see how structural lesions effect brain dynamics. In this hands-on session you will insert lesions into the connectome within the TVB graphical user interface (GUI). Afterwards, the modified connectome will be used for simulations and the resulting activity will be analysed using functional connectivity.
Difficulty level: Beginner
Duration: 31:22
Speaker: : Paul Triebkorn
Course:
In this lesson you will learn how to simulate seizure events and epilepsy in The Virtual Brain. We will look at the paper On the Nature of Seizure Dynamics, which describes a new local model called the Epileptor, and apply this same model in The Virtual Brain. This is part 1 of 2 in a series explaining how to use the Epileptor. In this part, we focus on setting up the parameters.
Difficulty level: Beginner
Duration: 4:44
Speaker: : Paul Triebkorn
This talk highlights a set of platform technologies, software, and data collections that close and shorten the feedback cycle in research.
Difficulty level: Beginner
Duration: 57:52
Speaker: : Satrajit Ghosh
Course:
This talk covers the Neuroimaging Informatics Tools and Resources Clearinghouse (NITRC), a free one-stop-shop collaboratory for science researchers that need resources such as neuroimaging analysis software, publicly available data sets, or computing power.
Difficulty level: Beginner
Duration: 1:00:10
Speaker: : David Kennedy
Course:
In this lecture, attendees will learn how Mutant Mouse Resource and Research Center (MMRRC) archives, cryopreserves, and distributes scientifically valuable genetically engineered mouse strains and mouse ES cell lines for the genetics and biomedical research community.
Difficulty level: Beginner
Duration: 43:38
Speaker: : Kent Lloyd
Course:
This lesson introduces the EEGLAB toolbox, as well as motivations for its use.
Difficulty level: Beginner
Duration: 15:32
Speaker: : Arnaud Delorme
Course:
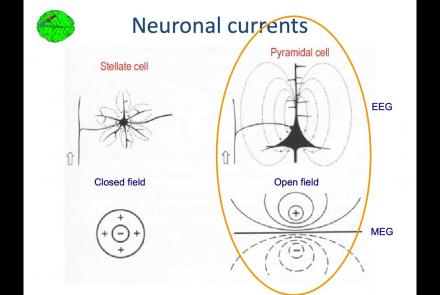
In this lesson, you will learn about the biological activity which generates and is measured by the EEG signal.
Difficulty level: Beginner
Duration: 6:53
Speaker: : Arnaud Delorme
Course:
This lesson goes over the characteristics of EEG signals when analyzed in source space (as opposed to sensor space).
Difficulty level: Beginner
Duration: 10:56
Speaker: : Arnaud Delorme
Course:
This lesson describes the development of EEGLAB as well as to what extent it is used by the research community.
Difficulty level: Beginner
Duration: 6:06
Speaker: : Arnaud Delorme
Course:
This lesson provides instruction as to how to build a processing pipeline in EEGLAB for a single participant.
Difficulty level: Beginner
Duration: 9:20
Speaker: :
Course:
Whereas the previous lesson of this course outlined how to build a processing pipeline for a single participant, this lesson discusses analysis pipelines for multiple participants simultaneously.
Difficulty level: Beginner
Duration: 10:55
Speaker: : Arnaud Delorme
Course:
In addition to outlining the motivations behind preprocessing EEG data in general, this lesson covers the first step in preprocessing data with EEGLAB, importing raw data.
Difficulty level: Beginner
Duration: 8:30
Speaker: : Arnaud Delorme
Topics
- Standards and Best Practices (1)
- Machine learning (1)
- Animal models (1)
- Assembly 2021 (6)
- Clinical neuroscience (1)
- Repositories and science gateways (1)
- Resources (3)
- General neuroscience (4)
- Phenome (1)
- General neuroinformatics (1)
- Computational neuroscience (20)
- Statistics (1)
- Computer Science (3)
- Genomics (21)
- Data science (14)
- Open science (20)
- Education (2)
- Publishing (1)