Lesson type
Difficulty level
This is the first of two workshops on reproducibility in science, during which participants are introduced to concepts of FAIR and open science. After discussing the definition of and need for FAIR science, participants are walked through tutorials on installing and using Github and Docker, the powerful, open-source tools for versioning and publishing code and software, respectively.
Difficulty level: Intermediate
Duration: 1:20:58
Speaker: : Erin Dickie and Sejal Patel
This lesson contains both a lecture and a tutorial component. The lecture (0:00-20:03 of YouTube video) discusses both the need for intersectional approaches in healthcare as well as the impact of neglecting intersectionality in patient populations. The lecture is followed by a practical tutorial in both Python and R on how to assess intersectional bias in datasets. Links to relevant code and data are found below.
Difficulty level: Beginner
Duration: 52:26
This is a hands-on tutorial on PLINK, the open source whole genome association analysis toolset. The aims of this tutorial are to teach users how to perform basic quality control on genetic datasets, as well as to identify and understand GWAS summary statistics.
Difficulty level: Intermediate
Duration: 1:27:18
Speaker: : Dan Felsky
This is a tutorial on using the open-source software PRSice to calculate a set of polygenic risk scores (PRS) for a study sample. Users will also learn how to read PRS into R, visualize distributions, and perform basic association analyses.
Difficulty level: Intermediate
Duration: 1:53:34
Speaker: : Dan Felsky
Course:
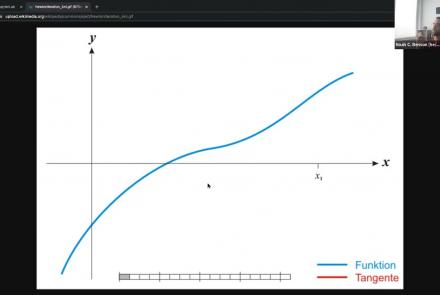
As a part of NeuroHackademy 2021, Noah Benson gives an introduction to Pytorch, one of the two most common software packages for deep learning applications to the neurosciences.
Difficulty level: Beginner
Duration: 00:50:40
Speaker: :
Course:
In this hands-on tutorial, Dr. Robert Guangyu Yang works through a number of coding exercises to see how RNNs can be easily used to study cognitive neuroscience questions, with a quick demonstration of how we can train and analyze RNNs on various cognitive neuroscience tasks. Familiarity of Python and basic knowledge of Pytorch are assumed.
Difficulty level: Beginner
Duration: 00:26:38
Speaker: :
Topics
- Bayesian networks (2)
- Cognitive neuroinformatics (1)
- Neuroimaging (15)
- Machine learning (3)
- Neuromorphic engineering (1)
- Standards and best practices (4)
- Tools (1)
- Animal models (1)
- Brain-hardware interfaces (1)
- Repositories and science gateways (1)
- General neuroscience (7)
- Computational neuroscience (21)
- Statistics (4)
- Computer Science (1)
- Genomics (5)
- Data science (10)
- (-) Open science (4)