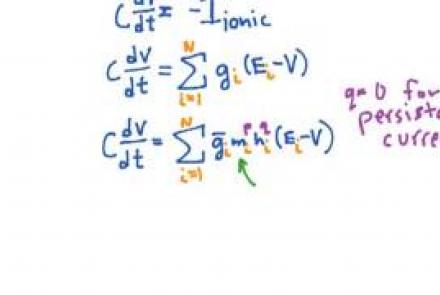Displaying 938 results
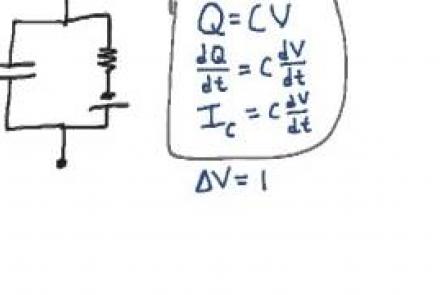
Explanation of the equivalent circuit model for a patch of passive neural membrane.
Difficulty level: Beginner
Duration: 9:22
Speaker: : Alex Williams
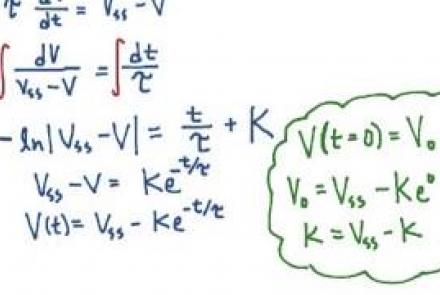
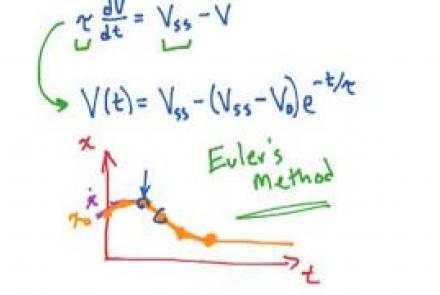
Solving the passive membrane equation
Difficulty level: Beginner
Duration: 4:04
Speaker: : Alex Williams
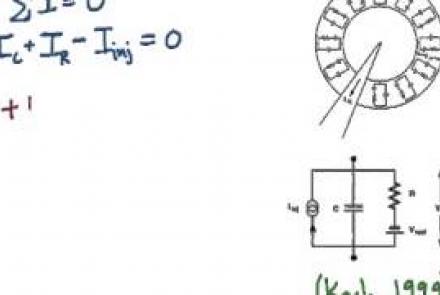
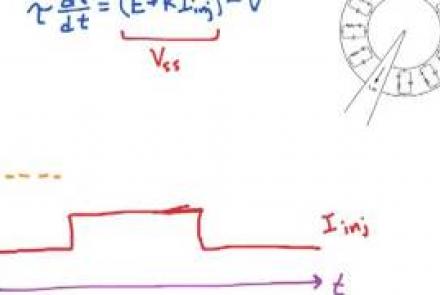
Injecting current into a passive membrane
Difficulty level: Beginner
Duration: 5:54
Speaker: : Alex Williams
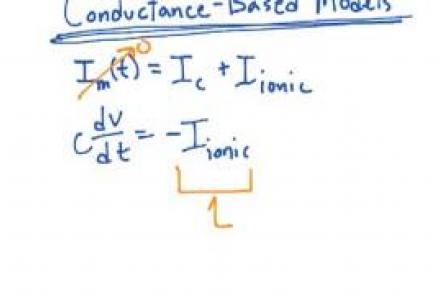
Explains the logic behind dealing with more complex currents by solving the membrane equation numerically.
Difficulty level: Beginner
Duration: 9:55
Speaker: : Alex Williams
Introducing voltage-dependent ion channels into the passive membrane
Difficulty level: Beginner
Duration: 8:32
Speaker: : Alex Williams
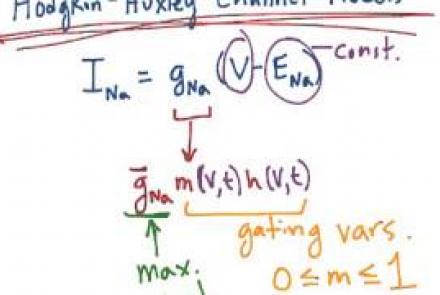
Introducing Hodgkin & Huxley's voltage dependent ion channel models, with emphasis on the sodium conductance
Difficulty level: Beginner
Duration: 10:32
Speaker: : Alex Williams
Introducing the classical Hodgkin & Huxley squid axon model with sodium and potassium conductances
Difficulty level: Beginner
Duration: 9:56
Speaker: : Alex Williams
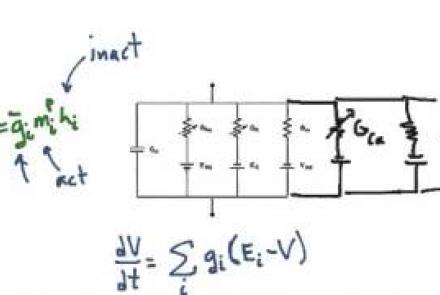
This lesson extends the conductance-based model equation to multiple neuronal compartments, taking more complex morphology into account.
Difficulty level: Beginner
Duration: 9:23
Speaker: : Alex Williams
Course:
This tutorial is part 1 of 2. It aims to provide viewers with an understanding of the fundamentals of R tool. Note: parts 1 and 2 of this tutorial are part of the same YouTube video; part 1 ends at 17:42.
Difficulty level: Beginner
Duration: 17:42
Speaker: : Edureka
Course:
This tutorial is part 2 of 2. It aims to provide viewers with an understanding of the fundamentals of R tool. Note: parts 1 and 2 of this tutorial are the same YouTube video. The portion related to this tutorial begins at 17:43.
Difficulty level: Beginner
Duration: 1:32:59
Speaker: : Edureka
The Allen Mouse Brain Atlas is a genome-wide, high-resolution atlas of gene expression throughout the adult mouse brain. This tutorial describes the basic search and navigation features of the Allen Mouse Brain Atlas.
Difficulty level: Beginner
Duration: 6:40
Speaker: : Allen Institute for Brain Science
The Allen Developing Mouse Brain Atlas is a detailed atlas of gene expression across mouse brain development. This tutorial describes the basic search and navigation features of the Allen Developing Mouse Brain Atlas.
Difficulty level: Beginner
Duration: 6:35
Speaker: : Unknown
This tutorial demonstrates how to use the differential search feature of the Allen Mouse Brain Atlas to find gene markers for different regions of the brain, as well as to visualize this gene expression in three-dimensional space. Differential search is also available for the Allen Developing Mouse Brain Atlas and the Allen Human Brain Atlas.
Difficulty level: Beginner
Duration: 6:31
Speaker: : Unknown
In this opening lesson, you will hear from the chair of the workshop (Neuroinformatics 2014 in Leiden, Netherlands), who gives an introduction and motivating argument underscoring the importance of collaboration in computational neuroscience.
Difficulty level: Beginner
Duration: 8:34
Speaker: : Angus Silver
This lesson gives an introduction to OpenWorm: an open-source project dedicated to creating a virtual C. elegans nematode in a computer.
Difficulty level: Intermediate
Duration: 23:26
Speaker: : Stephen Larson
The Open Source Brain (OSB) initiative (http://www.opensourcebrain.org) has been created to address the issues of poor accessibility, transparency, validation, and reuse of models in computational neuroscience.This lecture covers the aims of the Open Source Brain initiative, the current functionality of the website, and the range of models already available, and future plans for the project.
Difficulty level: Intermediate
Duration: 25:32
Speaker: : Padraig Gleeson
This lecture covers NeuronUnit, a library that builds upon SciUnit and integrates with several existing neuroinformatics resources to support validating single-neuron models using data gathered by neurophysiologists.
Difficulty level: Intermediate
Duration: 17:21
Speaker: : Richard Gerkin
This lesson provides an introduction to the NeuroElectro project, which aims to organize information on cellular neurophysiology.
Difficulty level: Intermediate
Duration: 17:41
Speaker: : Shreejoy Tripathy
In this lecture, the speaker demonstrates Neurokernel's module interfacing feature by using it to integrate independently developed models of olfactory and vision LPUs based upon experimentally obtained connectivity information.
Difficulty level: Intermediate
Duration: 29:56
Speaker: : Aurel A. Lazar