Course:
Longitudinal Online Research and Imaging System (LORIS) is a web-based data and project management software for neuroimaging research studies. It is an open source framework for storing and processing behavioural, clinical, neuroimaging and genetic data. LORIS also makes it easy to manage large datasets acquired over time in a longitudinal study, or at different locations in a large multi-site study.
Difficulty level: Beginner
Duration: 0:35
Speaker: : Samir Das
Course:
This lesson introduces the EEGLAB toolbox, as well as motivations for its use.
Difficulty level: Beginner
Duration: 15:32
Speaker: : Arnaud Delorme
Course:
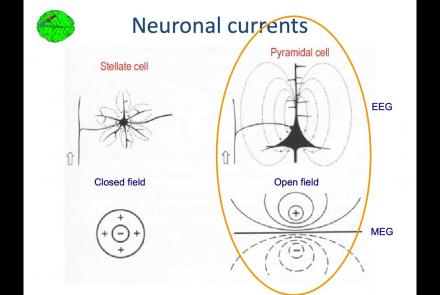
In this lesson, you will learn about the biological activity which generates and is measured by the EEG signal.
Difficulty level: Beginner
Duration: 6:53
Speaker: : Arnaud Delorme
Course:
This lesson goes over the characteristics of EEG signals when analyzed in source space (as opposed to sensor space).
Difficulty level: Beginner
Duration: 10:56
Speaker: : Arnaud Delorme
Course:
This lesson describes the development of EEGLAB as well as to what extent it is used by the research community.
Difficulty level: Beginner
Duration: 6:06
Speaker: : Arnaud Delorme
Course:
This lesson provides instruction as to how to build a processing pipeline in EEGLAB for a single participant.
Difficulty level: Beginner
Duration: 9:20
Speaker: :
Course:
Whereas the previous lesson of this course outlined how to build a processing pipeline for a single participant, this lesson discusses analysis pipelines for multiple participants simultaneously.
Difficulty level: Beginner
Duration: 10:55
Speaker: : Arnaud Delorme
Course:
In addition to outlining the motivations behind preprocessing EEG data in general, this lesson covers the first step in preprocessing data with EEGLAB, importing raw data.
Difficulty level: Beginner
Duration: 8:30
Speaker: : Arnaud Delorme
Course:
Continuing along the EEGLAB preprocessing pipeline, this tutorial walks users through how to import data events as well as EEG channel locations.
Difficulty level: Beginner
Duration: 11:53
Speaker: : Arnaud Delorme
Course:
This tutorial instructs users how to visually inspect partially pre-processed neuroimaging data in EEGLAB, specifically how to use the data browser to investigate specific channels, epochs, or events for removable artifacts, biological (e.g., eye blinks, muscle movements, heartbeat) or otherwise (e.g., corrupt channel, line noise).
Difficulty level: Beginner
Duration: 5:08
Speaker: : Arnaud Delorme
Course:
This tutorial provides instruction on how to use EEGLAB to further preprocess EEG datasets by identifying and discarding bad channels which, if left unaddressed, can corrupt and confound subsequent analysis steps.
Difficulty level: Beginner
Duration: 13:01
Speaker: : Arnaud Delorme
Course:
Users following this tutorial will learn how to identify and discard bad EEG data segments using the MATLAB toolbox EEGLAB.
Difficulty level: Beginner
Duration: 11:25
Speaker: : Arnaud Delorme
This lecture covers how to make modeling workflows FAIR by working through a practical example, dissecting the steps within the workflow, and detailing the tools and resources used at each step.
Difficulty level: Beginner
Duration: 15:14
Speaker: : Salvador Dura-Bernal
Course:
This lecture will provide an overview of the INCF Training Suite, a collection of tools that embraces the FAIR principles developed by members of the INCF Community. This will include an overview of TrainingSpace, Neurostars, and KnowledgeSpace.
Difficulty level: Beginner
Duration: 09:50
Speaker: : Mathew Abrams
Course:
This session will include presentations of infrastructure that embrace the FAIR principles developed by members of the INCF Community. This lecture provides an overview and demo of the Canadian Open Neuroscience Platform (CONP).
Difficulty level: Beginner
Duration: 14:02
Speaker: : Tristan Glatard & Samir Das
Course:
This session provides users with an introduction to tools and resources that facilitate the implementation of FAIR in their research.
Difficulty level: Beginner
Duration: 38:36
This lesson provides an overview of self-supervision as it relates to neural data tasks and the Mine Your Own vieW (MYOW) approach.
Difficulty level: Beginner
Duration: 25:50
Speaker: : Eva Dyer
Course:
This session will include presentations of infrastructure that embrace the FAIR principles developed by members of the INCF Community.
This lecture provides an overview of The Virtual Brain Simulation Platform.
Difficulty level: Beginner
Duration: 9:36
Speaker: : Petra Ritter
This video introduces the key principles for data organization and explains how you could make your data FAIR for data sharing on EBRAINS.
Difficulty level: Beginner
Duration: 10:54
Speaker: : Maaike van Swieten
This video explains what metadata is, why it is important, and how you can organize your metadata to increase the FAIRness of your data on EBRAINS.
Difficulty level: Beginner
Duration: 17:23
Speaker: : Ulrike Schlegel
Topics
- Standards and Best Practices (1)
- Machine learning (1)
- Animal models (1)
- (-) Assembly 2021 (6)
- Clinical neuroscience (1)
- Repositories and science gateways (1)
- Resources (3)
- General neuroscience (4)
- Phenome (1)
- General neuroinformatics (1)
- Computational neuroscience (20)
- Statistics (1)
- (-) Computer Science (3)
- Genomics (21)
- Data science (14)
- Open science (20)
- Education (2)
- Publishing (1)