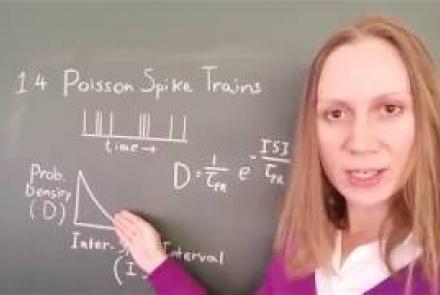Research Resource Identifiers (RRIDs) are ID numbers assigned to help researchers cite key resources (e.g., antibodies, model organisms, and software projects) in biomedical literature to improve the transparency of research methods.
Difficulty level: Beginner
Duration: 1:01:36
Speaker: : Maryann Martone
Course:
This lecture covers the rationale for developing the DAQCORD, a framework for the design, documentation, and reporting of data curation methods in order to advance the scientific rigour, reproducibility, and analysis of data.
Difficulty level: Intermediate
Duration: 17:08
Speaker: : Ari Ercole
Course:
The Mouse Phenome Database (MPD) provides access to primary experimental trait data, genotypic variation, protocols and analysis tools for mouse genetic studies. Data are contributed by investigators worldwide and represent a broad scope of phenotyping endpoints and disease-related traits in naïve mice and those exposed to drugs, environmental agents or other treatments. MPD ensures rigorous curation of phenotype data and supporting documentation using relevant ontologies and controlled vocabularies. As a repository of curated and integrated data, MPD provides a means to access/re-use baseline data, as well as allows users to identify sensitized backgrounds for making new mouse models with genome editing technologies, analyze trait co-inheritance, benchmark assays in their own laboratories, and many other research applications. MPD’s primary source of funding is NIDA. For this reason, a majority of MPD data is neuro- and behavior-related.
Difficulty level: Beginner
Duration: 55:36
Speaker: : Elissa Chesler
This lesson provides a brief introduction to the Computational Modeling of Neuronal Plasticity.
Difficulty level: Intermediate
Duration: 0:40
Speaker: : Florence I. Kleberg
In this lesson, you will be introducted to a type of neuronal model known as the leaky integrate-and-fire (LIF) model.
Difficulty level: Intermediate
Duration: 1:23
Speaker: : Florence I. Kleberg
This lesson goes over various potential inputs to neuronal synapses, loci of neural communication.
Difficulty level: Intermediate
Duration: 1:20
Speaker: : Florence I. Kleberg
This lesson describes the how and why behind implementing integration time steps as part of a neuronal model.
Difficulty level: Intermediate
Duration: 1:08
Speaker: : Florence I. Kleberg
In this lesson, you will learn about neural spike trains which can be characterized as having a Poisson distribution.
Difficulty level: Intermediate
Duration: 1:18
Speaker: : Florence I. Kleberg
This lesson covers spike-rate adaptation, the process by which a neuron's firing pattern decays to a low, steady-state frequency during the sustained encoding of a stimulus.
Difficulty level: Intermediate
Duration: 1:26
Speaker: : Florence I. Kleberg
This lesson provides a brief explanation of how to implement a neuron's refractory period in a computational model.
Difficulty level: Intermediate
Duration: 0:42
Speaker: : Florence I. Kleberg
In this lesson, you will learn a computational description of the process which tunes neuronal connectivity strength, spike-timing-dependent plasticity (STDP).
Difficulty level: Intermediate
Duration: 2:40
Speaker: : Florence I. Kleberg
This lesson reviews theoretical and mathematical descriptions of correlated spike trains.
Difficulty level: Intermediate
Duration: 2:54
Speaker: : Florence I. Kleberg
This lesson investigates the effect of correlated spike trains on spike-timing dependent plasticity (STDP).
Difficulty level: Intermediate
Duration: 1:43
Speaker: : Florence I. Kleberg
This lesson goes over synaptic normalisation, the homeostatic process by which groups of weighted inputs scale up or down their biases.
Difficulty level: Intermediate
Duration: 2:58
Speaker: : Florence I. Kleberg
In this lesson, you will learn about the intrinsic plasticity of single neurons.
Difficulty level: Intermediate
Duration: 2:08
Speaker: : Florence I. Kleberg
This lesson covers short-term facilitation, a process whereby a neuron's synaptic transmission is enhanced for a short (sub-second) period.
Difficulty level: Intermediate
Duration: 1:58
Speaker: : Florence I. Kleberg
This lesson describes short-term depression, a reduction of synaptic information transfer between neurons.
Difficulty level: Intermediate
Duration: 1:40
Speaker: : Florence I. Kleberg
This lesson briefly wraps up the course on Computational Modeling of Neuronal Plasticity.
Difficulty level: Intermediate
Duration: 0:37
Speaker: : Florence I. Kleberg
This lesson is the first of three hands-on tutorials as part of the workshop Research Workflows for Collaborative Neuroscience. This tutorial goes over how to visualize data with Scanpy, a scalable toolkit for analyzing single-cell gene expression.
Difficulty level: Intermediate
Duration: 25:26
Speaker: : David Feng & Frank Zappulla
This hands-on tutorial walks you through DataJoint platform, highlighting features and schema which can be used to build robost neuroscientific pipelines.
Difficulty level: Beginner
Duration: 26:06
Speaker: : Milagros Marin
Topics
- (-) Standards and Best Practices (2)
- Notebooks (2)
- Clinical neuroinformatics (2)
- Provenance (2)
- Artificial Intelligence (1)
- Digital brain atlasing (3)
- Neuroimaging (36)
- Neuromorphic engineering (1)
- Optogenetics (1)
- Standards and best practices (5)
- Tools (20)
- (-) Workflows (3)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (1)
- Assembly 2021 (1)
- Brain-hardware interfaces (13)
- Clinical neuroscience (1)
- Repositories and science gateways (1)
- Resources (1)
- General neuroscience
(12)
- (-) Phenome (1)
- Computational neuroscience (95)
- Statistics (5)
- Computer Science (4)
- Genomics (28)
- Data science (14)
- (-) Open science (18)
- Education (1)
- Neuroethics (1)