Lesson type
Difficulty level
In this lesson, you will learn in more detail about neuromorphic computing, that is, non-standard computational architectures that mimic some aspect of the way the brain works.
Difficulty level: Intermediate
Duration: 10:08
Speaker: : Dan Goodman
This video provides a very quick introduction to some of the neuromorphic sensing devices, and how they offer unique, low-power applications.
Difficulty level: Intermediate
Duration: 2:37
Speaker: : Dan Goodman
Course:
This lesson describes the principles underlying functional magnetic resonance imaging (fMRI), diffusion-weighted imaging (DWI), tractography, and parcellation. These tools and concepts are explained in a broader context of neural connectivity and mental health.
Difficulty level: Intermediate
Duration: 1:47:22
Speaker: : Erin Dickie and John Griffiths
Course:
This tutorial introduces pipelines and methods to compute brain connectomes from fMRI data. With corresponding code and repositories, participants can follow along and learn how to programmatically preprocess, curate, and analyze functional and structural brain data to produce connectivity matrices.
Difficulty level: Intermediate
Duration: 1:39:04
Speaker: : Erin Dickie and John Griffiths
This lesson introduces the practical exercises which accompany the previous lessons on animal and human connectomes in the brain and nervous system.
Difficulty level: Intermediate
Duration: 4:10
Speaker: : Dan Goodman
Course:
This lecture and tutorial focuses on measuring human functional brain networks, as well as how to account for inherent variability within those networks.
Difficulty level: Intermediate
Duration: 50:44
Speaker: : Caterina Gratton
This lecture goes into detailed description of how to process workflows in the virtual research environment (VRE), including approaches for standardization, metadata, containerization, and constructing and maintaining scientific pipelines.
Difficulty level: Intermediate
Duration: 1:03:55
Speaker: : Patrik Bey
Course:
This video will document the process of creating a pipeline rule for batch processing on brainlife.
Difficulty level: Intermediate
Duration: 0:57
Speaker: :
Course:
This video will document the process of launching a Jupyter Notebook for group-level analyses directly from brainlife.
Difficulty level: Intermediate
Duration: 0:53
Speaker: :
Course:
This lecture introduces you to the basics of the Amazon Web Services public cloud. It covers the fundamentals of cloud computing and goes through both the motivations and processes involved in moving your research computing to the cloud.
Difficulty level: Intermediate
Duration: 3:09:12
Speaker: : Amanda Tan & Ariel Rokem
This video gives a brief introduction to Neuro4ML's lessons on neuromorphic computing - the use of specialized hardware which either directly mimics brain function or is inspired by some aspect of the way the brain computes.
Difficulty level: Intermediate
Duration: 3:56
Speaker: : Dan Goodman
This lesson presents a simulation software for spatial model neurons and their networks designed primarily for GPUs.
Difficulty level: Intermediate
Duration: 21:15
Speaker: : Tadashi Yamazaki
The lecture covers a brief introduction to neuromorphic engineering, some of the neuromorphic networks that the speaker has developed, and their potential applications, particularly in machine learning.
Difficulty level: Intermediate
Duration: 19:57
Speaker: : Runchun Mark Wang
This lesson provides an overview of the current status in the field of neuroscientific ontologies, presenting examples of data organization and standards, particularly from neuroimaging and electrophysiology.
Difficulty level: Intermediate
Duration: 33:41
Speaker: : Yaroslav O. Halchenko
This tutorial provides instruction on how to simulate brain tumors with TVB (reproducing publication: Marinazzo et al. 2020 Neuroimage). This tutorial comprises a didactic video, jupyter notebooks, and full data set for the construction of virtual brains from patients and health controls.
Difficulty level: Intermediate
Duration: 10:01
The tutorial on modelling strokes in TVB includes a didactic video and jupyter notebooks (reproducing publication: Falcon et al. 2016 eNeuro).
Difficulty level: Intermediate
Duration: 7:43
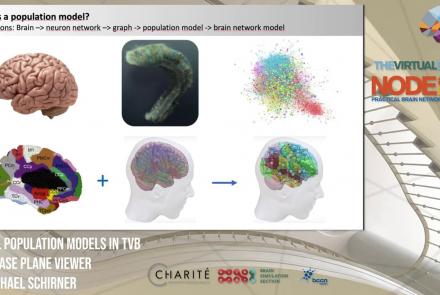
This lesson introduces population models and the phase plane, and is part of the The Virtual Brain (TVB) Node 10 Series, a 4-day workshop dedicated to learning about the full brain simulation platform TVB, as well as brain imaging, brain simulation, personalised brain models, and TVB use cases.
Difficulty level: Intermediate
Duration: 1:10:41
Speaker: : Michael Schirner
In this tutorial, you will learn how to run a typical TVB simulation.
Difficulty level: Intermediate
Duration: 1:29:13
Speaker: : Paul Triebkorn
This lesson introduces TVB-multi-scale extensions and other TVB tools which facilitate modeling and analyses of multi-scale data.
Difficulty level: Intermediate
Duration: 36:10
Speaker: : Dionysios Perdikis
This tutorial introduces The Virtual Mouse Brain (TVMB), walking users through the necessary steps for performing simulation operations on animal brain data.
Difficulty level: Intermediate
Duration: 42:43
Speaker: : Patrik Bey
Topics
- Artificial Intelligence (1)
- Notebooks (1)
- Provenance (1)
- DANDI archive (1)
- EBRAINS RI (6)
- Animal models (2)
- Brain-hardware interfaces (1)
- Clinical neuroscience (23)
- General neuroscience
(17)
- General neuroinformatics
(1)
- Computational neuroscience (80)
- Statistics (5)
- Computer Science (4)
- Genomics (7)
- Data science
(9)
- (-) Open science (5)
- Project management (1)
- Education (1)
- Neuroethics (5)