This is a hands-on tutorial on PLINK, the open source whole genome association analysis toolset. The aims of this tutorial are to teach users how to perform basic quality control on genetic datasets, as well as to identify and understand GWAS summary statistics.
Difficulty level: Intermediate
Duration: 1:27:18
Speaker: : Dan Felsky
This is an in-depth guide on EEG signals and their interaction within brain microcircuits. Participants are also shown techniques and software for simulating, analyzing, and visualizing these signals.
Difficulty level: Intermediate
Duration: 1:30:41
Speaker: : Frank Mazza
Course:
In this tutorial on simulating whole-brain activity using Python, participants can follow along using corresponding code and repositories, learning the basics of neural oscillatory dynamics, evoked responses and EEG signals, ultimately leading to the design of a network model of whole-brain anatomical connectivity.
Difficulty level: Intermediate
Duration: 1:16:10
Speaker: : John Griffiths
In this third and final hands-on tutorial from the Research Workflows for Collaborative Neuroscience workshop, you will learn about workflow orchestration using open source tools like DataJoint and Flyte.
Difficulty level: Intermediate
Duration: 22:36
Speaker: : Daniel Xenes
This lecture describes how to build research workflows, including a demonstrate using DataJoint Elements to build data pipelines.
Difficulty level: Intermediate
Duration: 47:00
Speaker: : Dimitri Yatsenko
This lesson introduces the practical exercises which accompany the previous lessons on animal and human connectomes in the brain and nervous system.
Difficulty level: Intermediate
Duration: 4:10
Speaker: : Dan Goodman
This lesson describes how DataLad allows you to track and mange both your data and analysis code, thereby facilitating reliable, reproducible, and shareable research.
Difficulty level: Intermediate
Duration: 59:34
Speaker: : Yaroslav O. Halchenko
This tutorial provides instruction on how to simulate brain tumors with TVB (reproducing publication: Marinazzo et al. 2020 Neuroimage). This tutorial comprises a didactic video, jupyter notebooks, and full data set for the construction of virtual brains from patients and health controls.
Difficulty level: Intermediate
Duration: 10:01
The tutorial on modelling strokes in TVB includes a didactic video and jupyter notebooks (reproducing publication: Falcon et al. 2016 eNeuro).
Difficulty level: Intermediate
Duration: 7:43
This tutorial covers the fundamentals of collaborating with Git and GitHub.
Difficulty level: Intermediate
Duration: 2:15:50
Speaker: : Elizabeth DuPre
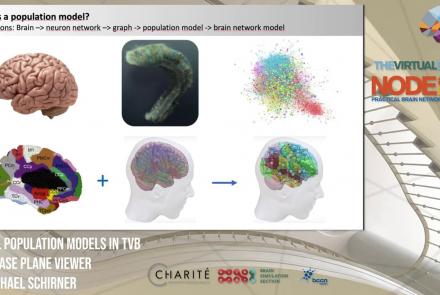
This lesson introduces population models and the phase plane, and is part of the The Virtual Brain (TVB) Node 10 Series, a 4-day workshop dedicated to learning about the full brain simulation platform TVB, as well as brain imaging, brain simulation, personalised brain models, and TVB use cases.
Difficulty level: Intermediate
Duration: 1:10:41
Speaker: : Michael Schirner
In this tutorial, you will learn how to run a typical TVB simulation.
Difficulty level: Intermediate
Duration: 1:29:13
Speaker: : Paul Triebkorn
This lesson introduces TVB-multi-scale extensions and other TVB tools which facilitate modeling and analyses of multi-scale data.
Difficulty level: Intermediate
Duration: 36:10
Speaker: : Dionysios Perdikis
This tutorial introduces The Virtual Mouse Brain (TVMB), walking users through the necessary steps for performing simulation operations on animal brain data.
Difficulty level: Intermediate
Duration: 42:43
Speaker: : Patrik Bey
In this tutorial, you will learn the necessary steps in modeling the brain of one of the most commonly studied animals among non-human primates, the macaque.
Difficulty level: Intermediate
Duration: 1:00:08
Speaker: : Julie Courtiol
This lecture delves into cortical (i.e., surface-based) brain simulations, as well as subcortical (i.e., deep brain) stimulations, covering the definitions, motivations, and implementations of both.
Difficulty level: Intermediate
Duration: 39:05
Speaker: : Jil Meier
This lecture provides an introduction to entropy in general, and multi-scale entropy (MSE) in particular, highlighting the potential clinical applications of the latter.
Difficulty level: Intermediate
Duration: 39:05
Speaker: : Jil Meier
This lecture gives an overview of how to prepare and preprocess neuroimaging (EEG/MEG) data for use in TVB.
Difficulty level: Intermediate
Duration: 1:40:52
Speaker: : Paul Triebkorn
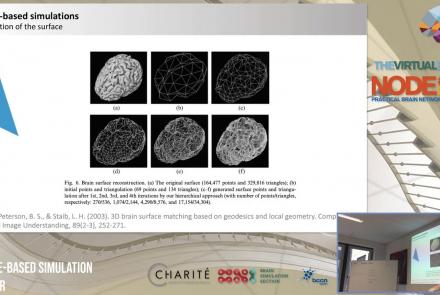
In this lecture, you will learn about various neuroinformatic resources which allow for 3D reconstruction of brain models.
Difficulty level: Intermediate
Duration: 1:36:57
Speaker: : Michael Schirner
Course:
This book was written with the goal of introducing researchers and students in a variety of research fields to the intersection of data science and neuroimaging. This book reflects our own experience of doing research at the intersection of data science and neuroimaging and it is based on our experience working with students and collaborators who come from a variety of backgrounds and have a variety of reasons for wanting to use data science approaches in their work. The tools and ideas that we chose to write about are all tools and ideas that we have used in some way in our own research. Many of them are tools that we use on a daily basis in our work. This was important to us for a few reasons: the first is that we want to teach people things that we ourselves find useful. Second, it allowed us to write the book with a focus on solving specific analysis tasks. For example, in many of the chapters you will see that we walk you through ideas while implementing them in code, and with data. We believe that this is a good way to learn about data analysis, because it provides a connecting thread from scientific questions through the data and its representation to implementing specific answers to these questions. Finally, we find these ideas compelling and fruitful. That’s why we were drawn to them in the first place. We hope that our enthusiasm about the ideas and tools described in this book will be infectious enough to convince the readers of their value.
Difficulty level: Intermediate
Duration:
Speaker: :
Topics
- Clinical neuroinformatics (5)
- Standards and Best Practices (1)
- Machine learning (6)
- Neuroimaging (5)
- EBRAINS RI (2)
- Neuromorphic engineering (1)
- Standards and best practices (8)
- (-) Tools (11)
- Workflows (4)
- Animal models (1)
- Brain-hardware interfaces (1)
- Clinical neuroscience (2)
- General neuroscience (3)
- Computational neuroscience (26)
- Statistics (2)
- (-) Computer Science (2)
- Genomics (5)
- Data science (4)
- Open science (4)
- Project management (1)