This is a hands-on tutorial on PLINK, the open source whole genome association analysis toolset. The aims of this tutorial are to teach users how to perform basic quality control on genetic datasets, as well as to identify and understand GWAS summary statistics.
Difficulty level: Intermediate
Duration: 1:27:18
Speaker: : Dan Felsky
In this workshop talk, you will receive a tour of the Code Ocean ScienceOps Platform, a centralized cloud workspace for all teams.
Difficulty level: Beginner
Duration: 10:24
Speaker: : Frank Zappulla
This talk describes approaches to maintaining integrated workflows and data management schema, taking advantage of the many open source, collaborative platforms already existing.
Difficulty level: Beginner
Duration: 15:15
Speaker: : Erik C. Johnson
In this third and final hands-on tutorial from the Research Workflows for Collaborative Neuroscience workshop, you will learn about workflow orchestration using open source tools like DataJoint and Flyte.
Difficulty level: Intermediate
Duration: 22:36
Speaker: : Daniel Xenes
This lesson provides an introduction to the DataLad, a free and open source distributed data management system that keeps track of your data, creates structure, ensures reproducibility, supports collaboration, and integrates with widely used data infrastructure.
Difficulty level: Beginner
Duration: 22:56
Speaker: : Michał Szczepanik
This lesson introduces several open science tools like Docker and Apptainer which can be used to develop portable and reproducible software environments.
Difficulty level: Beginner
Duration: 17:22
Speaker: : Joanes Grandjean
This lecture provides a detailed description of how to incorporate HED annotation into your neuroimaging data pipeline.
Difficulty level: Beginner
Duration: 33:36
Speaker: : Dung Truong
This talk covers the differences between applying HED annotation to fMRI datasets versus other neuroimaging practices, and also introduces an analysis pipeline using HED tags.
Difficulty level: Beginner
Duration: 22:52
Speaker: : Monique Denissen
This lecture describes how to build research workflows, including a demonstrate using DataJoint Elements to build data pipelines.
Difficulty level: Intermediate
Duration: 47:00
Speaker: : Dimitri Yatsenko
Following the previous lesson on neuronal structure, this lesson discusses neuronal function, particularly focusing on spike triggering and propogation.
Difficulty level: Intermediate
Duration: 6:58
Speaker: : Marcus Ghosh
Course:
This video will teach you the basics of navigating the Open Science Framework and creating your first projects.
Difficulty level: Beginner
Duration: 2:11
Speaker: :
Course:
This webinar walks you through the basics of creating an OSF project, structuring it to fit your research needs, adding collaborators, and tying your favorite online tools into your project structure.
Difficulty level: Beginner
Duration: 55:02
Speaker: : Ian Sullivan
Course:
This webinar will introduce how to use the Open Science Framework (OSF) in a classroom setting.
Difficulty level: Beginner
Duration: 32:01
Speaker: : April Clyburne-Sherin
Course:
This lesson provides instruction on how to organize related projects with OSF features such as links, forks, and templates.
Difficulty level: Beginner
Duration: 51:14
Speaker: : Ian Sullivan
This webinar will introduce the integration of JASP Statistical Software with the Open Science Framework (OSF).
Difficulty level: Beginner
Duration: 30:56
Speaker: : Alexander Etz
Course:
This lesson describes the value of version control, as well as how to do so with your own files and data on OSF.
Difficulty level: Beginner
Duration: 22:07
Speaker: : Courtney Soderberg
Course:
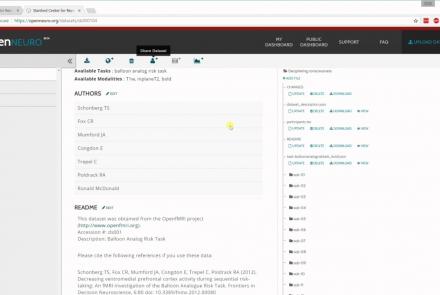
In this tutorial, you will learn the basic features of uploading and versioning your data within OpenNeuro.org.
Difficulty level: Beginner
Duration: 5:36
Speaker: : OpenNeuro
Course:
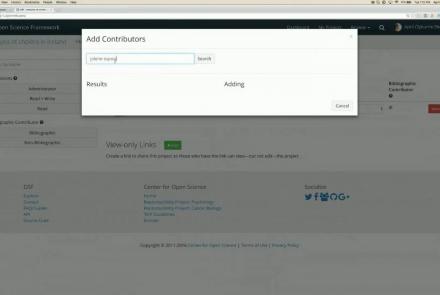
This tutorial shows how to share your data in OpenNeuro.org.
Difficulty level: Beginner
Duration: 1:22
Speaker: : OpenNeuro
Course:
Following the previous two tutorials on uploading and sharing data with OpenNeuro.org, this tutorial briefly covers how to run various analyses on your datasets.
Difficulty level: Beginner
Duration: 2:26
Speaker: : OpenNeuro
This lesson introduces the practical usage of The Virtual Brain (TVB) in its graphical user interface and via python scripts. In the graphical user interface, you are guided through its data repository, simulator, phase plane exploration tool, connectivity editor, stimulus generator, and the provided analyses. The implemented iPython notebooks of TVB are presented, and since they are public, can be used for further exploration of TVB.
Difficulty level: Beginner
Duration: 1:12:24
Speaker: : Paul Triebkorn
Topics
- Standards and Best Practices (2)
- Machine learning (2)
- Animal models (1)
- Assembly 2021 (6)
- Clinical neuroscience (3)
- Repositories and science gateways (1)
- Resources (3)
- (-) General neuroscience (5)
- Phenome (1)
- General neuroinformatics (1)
- (-) Computational neuroscience (42)
- Statistics (2)
- Computer Science (4)
- Genomics (26)
- Data science (17)
- Open science (24)
- Project management (1)
- Education (2)
- Publishing (1)