Course:
This lesson introduces the EEGLAB toolbox, as well as motivations for its use.
Difficulty level: Beginner
Duration: 15:32
Speaker: : Arnaud Delorme
Course:
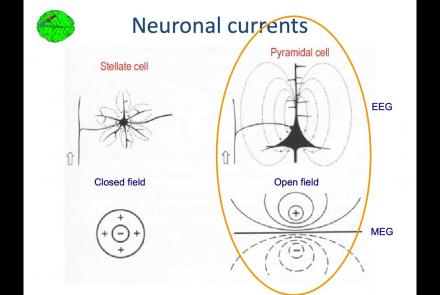
In this lesson, you will learn about the biological activity which generates and is measured by the EEG signal.
Difficulty level: Beginner
Duration: 6:53
Speaker: : Arnaud Delorme
Course:
This lesson goes over the characteristics of EEG signals when analyzed in source space (as opposed to sensor space).
Difficulty level: Beginner
Duration: 10:56
Speaker: : Arnaud Delorme
Course:
This lesson describes the development of EEGLAB as well as to what extent it is used by the research community.
Difficulty level: Beginner
Duration: 6:06
Speaker: : Arnaud Delorme
Course:
This lesson provides instruction as to how to build a processing pipeline in EEGLAB for a single participant.
Difficulty level: Beginner
Duration: 9:20
Speaker: :
Course:
Whereas the previous lesson of this course outlined how to build a processing pipeline for a single participant, this lesson discusses analysis pipelines for multiple participants simultaneously.
Difficulty level: Beginner
Duration: 10:55
Speaker: : Arnaud Delorme
Course:
In addition to outlining the motivations behind preprocessing EEG data in general, this lesson covers the first step in preprocessing data with EEGLAB, importing raw data.
Difficulty level: Beginner
Duration: 8:30
Speaker: : Arnaud Delorme
Course:
Continuing along the EEGLAB preprocessing pipeline, this tutorial walks users through how to import data events as well as EEG channel locations.
Difficulty level: Beginner
Duration: 11:53
Speaker: : Arnaud Delorme
Course:
This tutorial instructs users how to visually inspect partially pre-processed neuroimaging data in EEGLAB, specifically how to use the data browser to investigate specific channels, epochs, or events for removable artifacts, biological (e.g., eye blinks, muscle movements, heartbeat) or otherwise (e.g., corrupt channel, line noise).
Difficulty level: Beginner
Duration: 5:08
Speaker: : Arnaud Delorme
Course:
This tutorial provides instruction on how to use EEGLAB to further preprocess EEG datasets by identifying and discarding bad channels which, if left unaddressed, can corrupt and confound subsequent analysis steps.
Difficulty level: Beginner
Duration: 13:01
Speaker: : Arnaud Delorme
Course:
Users following this tutorial will learn how to identify and discard bad EEG data segments using the MATLAB toolbox EEGLAB.
Difficulty level: Beginner
Duration: 11:25
Speaker: : Arnaud Delorme
This lecture gives an overview of how to prepare and preprocess neuroimaging (EEG/MEG) data for use in TVB.
Difficulty level: Intermediate
Duration: 1:40:52
Speaker: : Paul Triebkorn
Course:
This lesson describes the principles underlying functional magnetic resonance imaging (fMRI), diffusion-weighted imaging (DWI), tractography, and parcellation. These tools and concepts are explained in a broader context of neural connectivity and mental health.
Difficulty level: Intermediate
Duration: 1:47:22
Speaker: : Erin Dickie and John Griffiths
This talk covers the differences between applying HED annotation to fMRI datasets versus other neuroimaging practices, and also introduces an analysis pipeline using HED tags.
Difficulty level: Beginner
Duration: 22:52
Speaker: : Monique Denissen
This lesson explores how researchers try to understand neural networks, particularly in the case of observing neural activity.
Difficulty level: Intermediate
Duration: 8:20
Speaker: : Marcus Ghosh
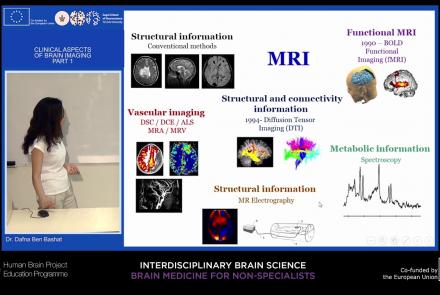
This lecture will provide an overview of neuroimaging techniques and their clinical applications.
Difficulty level: Beginner
Duration: 45:29
Speaker: : Dafna Ben Bashat
Course:
Longitudinal Online Research and Imaging System (LORIS) is a web-based data and project management software for neuroimaging research studies. It is an open source framework for storing and processing behavioural, clinical, neuroimaging and genetic data. LORIS also makes it easy to manage large datasets acquired over time in a longitudinal study, or at different locations in a large multi-site study.
Difficulty level: Beginner
Duration: 0:35
Speaker: : Samir Das
Course:
This talk covers the Neuroimaging Informatics Tools and Resources Clearinghouse (NITRC), a free one-stop-shop collaboratory for science researchers that need resources such as neuroimaging analysis software, publicly available data sets, or computing power.
Difficulty level: Beginner
Duration: 1:00:10
Speaker: : David Kennedy
This lecture provides an introduction to the Brain Imaging Data Structure (BIDS), a standard for organizing human neuroimaging datasets.
Difficulty level: Intermediate
Duration: 56:49
Speaker: : Chris Gorgolewski
Course:
This lecture presents an overview of functional brain parcellations, as well as a set of tutorials on bootstrap agregation of stable clusters (BASC) for fMRI brain parcellation.
Difficulty level: Advanced
Duration: 50:28
Speaker: : Pierre Bellec
Topics
- Philosophy of Science (5)
- Artificial Intelligence (4)
- Animal models (3)
- Assembly 2021 (27)
- Brain-hardware interfaces (2)
- Clinical neuroscience (14)
- International Brain Initiative (2)
- Repositories and science gateways (5)
- Resources (6)
- General neuroscience
(18)
- Cognitive Science (7)
- Cell signaling (3)
- Brain networks (6)
- Glia (1)
- Electrophysiology (14)
- Learning and memory (4)
- Neuroanatomy (3)
- Neurobiology (12)
- Neurodegeneration (1)
- Neuroimmunology (1)
- Neural networks (11)
- Neurophysiology (6)
- Neuropharmacology (1)
- Neuronal plasticity (1)
- Synaptic plasticity (1)
- Phenome (1)
- General neuroinformatics
(13)
- Computational neuroscience (95)
- Statistics (4)
- Computer Science (7)
- Genomics (29)
- Data science (20)
- Open science (34)
- Project management (4)
- Education (2)
- Publishing (1)
- (-) Neuroethics (6)