This lecture describes how to build research workflows, including a demonstrate using DataJoint Elements to build data pipelines.
Difficulty level: Intermediate
Duration: 47:00
Speaker: : Dimitri Yatsenko
This lesson provides an introduction to the Symposium on Science Management at the Canadian Association for Neuroscience 2019 Meeting.
Difficulty level: Beginner
Duration: 9:52
Speaker: : Randy McIntosh
This lesson gives a primer to project management in a scientific context, with a particular neuroinformatic case study.
Difficulty level: Beginner
Duration: 19:06
Speaker: : Kelly Shen
In this lesson, you will hear about the current challenges regarding data management, as well as policies and resources aimed to address them.
Difficulty level: Beginner
Duration: 18:13
Speaker: : Mojib Javadi
This lesson covers "Knowledge Translation", the activities involved in moving research from the laboratory, the research journal, and the academic conference into the hands of people and organizations who can put it to practical use.
Difficulty level: Beginner
Duration: 15:05
Speaker: : Jordan Antflick
In this lesson, you will hear about the various methods developed and employed in managing performance.
Difficulty level: Beginner
Duration: 12:57
Speaker: : Christa Studzinski
This lesson provides an overview of how to manage relationships in a research context, while highlighting the need for effective communication at various levels.
Difficulty level: Beginner
Duration:
Speaker: : Helena Ledmyr
In this lesson you will hear a panel discussion which hosts experts in the field whom have extensive experience with management in a science setting.
Difficulty level: Beginner
Duration: 54:38
Speaker: :
Learn how to create a standard extracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 23:10
Speaker: : Ryan Ly
Learn how to create a standard calcium imaging dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 31:04
Speaker: : Ryan Ly
In this tutorial, you will learn how to create a standard intracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 20:23
Speaker: : Pamela Baker
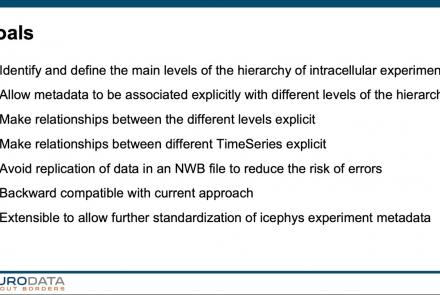
In this tutorial, you will learn how to use the icephys-metadata extension to enter meta-data detailing your experimental paradigm.
Difficulty level: Intermediate
Duration: 27:18
Speaker: : Oliver Ruebel
This lesson provides instructions on how to build and share extensions in NWB.
Difficulty level: Advanced
Duration: 20:29
Speaker: : Ryan Ly
Learn how to build custom APIs for extension.
Difficulty level: Advanced
Duration: 25:40
Speaker: : Andrew Tritt
This lesson provides instruction on advanced writing strategies in HDF5 that are accessible through PyNWB.
Difficulty level: Advanced
Duration: 23:00
Speaker: : Oliver Ruebel
In this tutorial, users learn how to create a standard extracellular electrophysiology dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 45:46
Speaker: : Ben Dichter
Learn how to create a standard calcium imaging dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 39:10
Speaker: : Ben Dichter
Learn how to create a standard intracellular electrophysiology dataset in NWB.
Difficulty level: Intermediate
Duration: 20:22
Speaker: : Pamela Baker
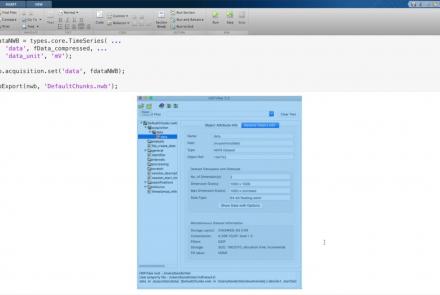
This lesson provides a tutorial on how to handle writing very large data in MatNWB.
Difficulty level: Advanced
Duration: 16:18
Speaker: : Ben Dichter
This lesson gives an overview of the Brainstorm package for analyzing extracellular electrophysiology, including preprocessing, spike sorting, trial alignment, and spectrotemporal decomposition.
Difficulty level: Intermediate
Duration: 47:47
Speaker: : Konstantinos Nasiotis
Topics
- Artificial Intelligence (7)
- Philosophy of Science (5)
- Provenance (3)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (8)
- Assembly 2021 (29)
- Brain-hardware interfaces (14)
- Clinical neuroscience (40)
- International Brain Initiative (2)
- Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(62)
- Neuroscience (11)
- Cognitive Science (7)
- Cell signaling (6)
- Brain networks (11)
- Glia (1)
- Electrophysiology (41)
- Learning and memory (5)
- Neuroanatomy (24)
- Neurobiology (16)
- Neurodegeneration (1)
- (-) Neuroimmunology (1)
- Neural networks (15)
- (-) Neurophysiology (27)
- (-) Neuropharmacology (2)
- Neuronal plasticity (16)
- Synaptic plasticity (4)
- (-) Visual system (12)
- Phenome (1)
- General neuroinformatics
(27)
- Computational neuroscience (278)
- Statistics (7)
- Computer Science (20)
- Genomics (33)
- Data science
(33)
- Open science (58)
- (-) Project management (8)
- Education (4)
- Publishing (4)
- Neuroethics (42)