This talk presents state-of-the-art methods for ensuring data privacy with a particular focus on medical data sharing across multiple organizations.
Difficulty level: Intermediate
Duration: 22:49
Speaker: : Barbara Carminati
This lecture talks about the usage of knowledge graphs in hospitals and related challenges of semantic interoperability.
Difficulty level: Intermediate
Duration: 24:32
Speaker: : Cristophe Gaudet-Blavignac
This lecture gives an overview of how to prepare and preprocess neuroimaging (EEG/MEG) data for use in TVB.
Difficulty level: Intermediate
Duration: 1:40:52
Speaker: : Paul Triebkorn
Course:
This lesson is a general overview of overarching concepts in neuroinformatics research, with a particular focus on clinical approaches to defining, measuring, studying, diagnosing, and treating various brain disorders. Also described are the complex, multi-level nature of brain disorders and the data associated with them, from genes and individual cells up to cortical microcircuits and whole-brain network dynamics. Given the heterogeneity of brain disorders and their underlying mechanisms, this lesson lays out a case for multiscale neuroscience data integration.
Difficulty level: Intermediate
Duration: 1:09:33
Speaker: : Sean Hill
This lesson breaks down the principles of Bayesian inference and how it relates to cognitive processes and functions like learning and perception. It is then explained how cognitive models can be built using Bayesian statistics in order to investigate how our brains interface with their environment.
This lesson corresponds to slides 1-64 in the PDF below.
Difficulty level: Intermediate
Duration: 1:28:14
Speaker: : Andreea Diaconescu
Course:
This lecture and tutorial focuses on measuring human functional brain networks, as well as how to account for inherent variability within those networks.
Difficulty level: Intermediate
Duration: 50:44
Speaker: : Caterina Gratton
This lesson continues from part one of the lecture Ontologies, Databases, and Standards, diving deeper into a description of ontologies and knowledg graphs.
Difficulty level: Intermediate
Duration: 50:18
Speaker: : Jeff Grethe
This lecture focuses on ontologies for clinical neurosciences.
Difficulty level: Intermediate
Duration: 21:54
Speaker: : Martin Hofmann-Apitius
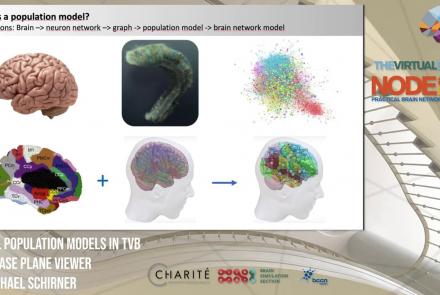
This lesson introduces population models and the phase plane, and is part of the The Virtual Brain (TVB) Node 10 Series, a 4-day workshop dedicated to learning about the full brain simulation platform TVB, as well as brain imaging, brain simulation, personalised brain models, and TVB use cases.
Difficulty level: Intermediate
Duration: 1:10:41
Speaker: : Michael Schirner
This lesson introduces TVB-multi-scale extensions and other TVB tools which facilitate modeling and analyses of multi-scale data.
Difficulty level: Intermediate
Duration: 36:10
Speaker: : Dionysios Perdikis
This lecture delves into cortical (i.e., surface-based) brain simulations, as well as subcortical (i.e., deep brain) stimulations, covering the definitions, motivations, and implementations of both.
Difficulty level: Intermediate
Duration: 39:05
Speaker: : Jil Meier
This lecture provides an introduction to entropy in general, and multi-scale entropy (MSE) in particular, highlighting the potential clinical applications of the latter.
Difficulty level: Intermediate
Duration: 39:05
Speaker: : Jil Meier
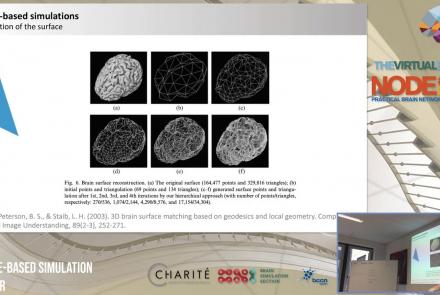
In this lecture, you will learn about various neuroinformatic resources which allow for 3D reconstruction of brain models.
Difficulty level: Intermediate
Duration: 1:36:57
Speaker: : Michael Schirner
This lecture discusses the the importance and need for data sharing in clinical neuroscience.
Difficulty level: Intermediate
Duration: 25:22
Speaker: : Thomas Berger
This lecture presents the Medical Informatic Platform's data federation for Traumatic Brain Injury.
Difficulty level: Intermediate
Duration: 25:55
Speaker: : Stefano Finazzi
This lecture gives insights into the Medical Informatics Platform's current and future data privacy model.
Difficulty level: Intermediate
Duration: 17:29
Speaker: : Yannis Ioannidis
This lecture explains the concept of federated analysis in the context of medical data, associated challenges. The lecture also presents an example of hospital federations via the Medical Informatics Platform.
Difficulty level: Intermediate
Duration: 19:15
Speaker: : Yannis Ioannidis
This lecture gives an overview on the European Health Dataspace.
Difficulty level: Intermediate
Duration: 26:33
Speaker: : Licino Kustra Mano
This lecture presents the Medical Informatics Platform's data federation in epilepsy.
Difficulty level: Intermediate
Duration: 27:09
Speaker: : Philippe Ryvlin
This lecture presents the Medical Informatics Platform's data federation in epilepsy.
Difficulty level: Intermediate
Duration: 26:51
Speaker: : Pawel Swieboda
Topics
- Artificial Intelligence (1)
- Provenance (1)
- EBRAINS RI (6)
- Brain-hardware interfaces (1)
- (-) Clinical neuroscience (20)
- General neuroscience (8)
- General neuroinformatics (1)
- Computational neuroscience (36)
- Statistics (2)
- (-) Computer Science (2)
- Genomics (2)
- Data science (7)
- Open science (1)
- Project management (1)
- Neuroethics (3)