Course:
This tutorial demonstrates how to work with neuronal data using MATLAB, including actional potentials and spike counts, orientation tuing curves in visual cortex, and spatial maps of firing rates.
Difficulty level: Intermediate
Duration: 5:17
Speaker: : Mike X. Cohen
Course:
This lesson instructs users on how to import electrophysiological neural data into MATLAB, as well as how to convert spikes to a data matrix.
Difficulty level: Intermediate
Duration: 11:37
Speaker: : Mike X. Cohen
This is a hands-on tutorial on PLINK, the open source whole genome association analysis toolset. The aims of this tutorial are to teach users how to perform basic quality control on genetic datasets, as well as to identify and understand GWAS summary statistics.
Difficulty level: Intermediate
Duration: 1:27:18
Speaker: : Dan Felsky
This is a tutorial on using the open-source software PRSice to calculate a set of polygenic risk scores (PRS) for a study sample. Users will also learn how to read PRS into R, visualize distributions, and perform basic association analyses.
Difficulty level: Intermediate
Duration: 1:53:34
Speaker: : Dan Felsky
Course:
This is an introductory lecture on whole-brain modelling, delving into the various spatial scales of neuroscience, neural population models, and whole-brain modelling. Additionally, the clinical applications of building and testing such models are characterized.
Difficulty level: Intermediate
Duration: 1:24:44
Speaker: : John Griffiths
This lecture aims to help researchers, students, and health care professionals understand the place for neuroinformatics in the patient journey using the exemplar of an epilepsy patient.
Difficulty level: Intermediate
Duration: 1:32:53
Speaker: : Randy Gollub & Prantik Kundu
Following the previous lesson on neuronal structure, this lesson discusses neuronal function, particularly focusing on spike triggering and propogation.
Difficulty level: Intermediate
Duration: 6:58
Speaker: : Marcus Ghosh
This lesson describes spike timing-dependent plasticity (STDP), a biological process that adjusts the strength of connections between neurons in the brain, and how one can implement or mimic this process in a computational model. You will also find links for practical exercises at the bottom of this page.
Difficulty level: Intermediate
Duration: 12:50
Speaker: : Dan Goodman
As the previous lesson of this course described how researchers acquire neural data, this lesson will discuss how to go about interpreting and analysing the data.
Difficulty level: Intermediate
Duration: 9:24
Speaker: : Marcus Ghosh
In this lesson you will learn about the motivation behind manipulating neural activity, and what forms that may take in various experimental designs.
Difficulty level: Intermediate
Duration: 8:42
Speaker: : Marcus Ghosh
In this lesson, you will learn about one particular aspect of decision making: reaction times. In other words, how long does it take to take a decision based on a stream of information arriving continuously over time?
Difficulty level: Intermediate
Duration: 6:01
Speaker: : Dan Goodman
In this lesson, you will hear about some of the open issues in the field of neuroscience, as well as a discussion about whether neuroscience works, and how can we know?
Difficulty level: Intermediate
Duration: 6:54
Speaker: : Marcus Ghosh
This lesson discusses a gripping neuroscientific question: why have neurons developed the discrete action potential, or spike, as a principle method of communication?
Difficulty level: Intermediate
Duration: 9:34
Speaker: : Dan Goodman
This tutorial covers the fundamentals of collaborating with Git and GitHub.
Difficulty level: Intermediate
Duration: 2:15:50
Speaker: : Elizabeth DuPre
Course:
This lesson provides an overview of Jupyter notebooks, Jupyter lab, and Binder, as well as their applications within the field of neuroimaging, particularly when it comes to the writing phase of your research.
Difficulty level: Intermediate
Duration: 50:28
Speaker: : Elizabeth DuPre
This lecture covers how you can make your data public through EBRAINS. This talk focuses on the ethical considerations for sharing data, the requirements that are imposed by various regulations, particularly for sharing human data. The lecture also includes a discussion of how EBRAINS designs its services to deal with the ethical and regulatory aspects of sharing these kinds of data.
Difficulty level: Intermediate
Duration: 16:15
Speaker: : Maaike van Swieten and Jan Bjaalie
This lecture discusses differential privacy and synthetic data in the context of medical data sharing in clinical neurosciences.
Difficulty level: Intermediate
Duration: 20:26
Speaker: : Minos Garofalakis
This talk presents state-of-the-art methods for ensuring data privacy with a particular focus on medical data sharing across multiple organizations.
Difficulty level: Intermediate
Duration: 22:49
Speaker: : Barbara Carminati
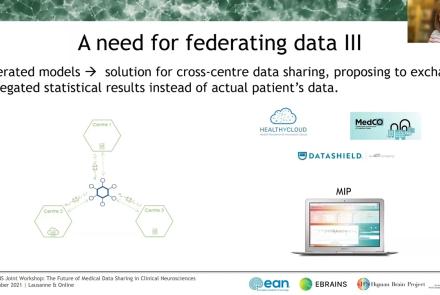
This talks presents ethics requirements of the Medical Informatics Platform, a data sharing platform for medical data using data federation mechanisms. The talk presents how the Medical Informatics Platform (MIP) works and which ethical requirements need to be considered when working with federated data.
Difficulty level: Intermediate
Duration: 16:25
Speaker: : Erika Borcel
This lecture talks about the usage of knowledge graphs in hospitals and related challenges of semantic interoperability.
Difficulty level: Intermediate
Duration: 24:32
Speaker: : Cristophe Gaudet-Blavignac
Topics
- Artificial Intelligence (1)
- Provenance (1)
- EBRAINS RI (6)
- Brain-hardware interfaces (1)
- Clinical neuroscience (20)
- (-) General neuroscience (9)
- General neuroinformatics (1)
- Computational neuroscience (45)
- Statistics (3)
- (-) Computer Science (3)
- Genomics (7)
- Data science (8)
- (-) Open science (4)
- Project management (1)
- (-) Neuroethics (3)