Course:
This lesson is a general overview of overarching concepts in neuroinformatics research, with a particular focus on clinical approaches to defining, measuring, studying, diagnosing, and treating various brain disorders. Also described are the complex, multi-level nature of brain disorders and the data associated with them, from genes and individual cells up to cortical microcircuits and whole-brain network dynamics. Given the heterogeneity of brain disorders and their underlying mechanisms, this lesson lays out a case for multiscale neuroscience data integration.
Difficulty level: Intermediate
Duration: 1:09:33
Speaker: : Sean Hill
Course:
In this tutorial on simulating whole-brain activity using Python, participants can follow along using corresponding code and repositories, learning the basics of neural oscillatory dynamics, evoked responses and EEG signals, ultimately leading to the design of a network model of whole-brain anatomical connectivity.
Difficulty level: Intermediate
Duration: 1:16:10
Speaker: : John Griffiths
This lesson breaks down the principles of Bayesian inference and how it relates to cognitive processes and functions like learning and perception. It is then explained how cognitive models can be built using Bayesian statistics in order to investigate how our brains interface with their environment.
This lesson corresponds to slides 1-64 in the PDF below.
Difficulty level: Intermediate
Duration: 1:28:14
Speaker: : Andreea Diaconescu
Whereas the previous two lessons described the biophysical and signalling properties of individual neurons, this lesson describes properties of those units when part of larger networks.
Difficulty level: Intermediate
Duration: 6:00
Speaker: : Marcus Ghosh
This lesson goes over some examples of how machine learners and computational neuroscientists go about designing and building neural network models inspired by biological brain systems.
Difficulty level: Intermediate
Duration: 12:52
Speaker: : Dan Goodman
Course:
This lecture and tutorial focuses on measuring human functional brain networks, as well as how to account for inherent variability within those networks.
Difficulty level: Intermediate
Duration: 50:44
Speaker: : Caterina Gratton
This is the Introductory Module to the Deep Learning Course at CDS, a course that covered the latest techniques in deep learning and representation learning, focusing on supervised and unsupervised deep learning, embedding methods, metric learning, convolutional and recurrent nets, with applications to computer vision, natural language understanding, and speech recognition.
Difficulty level: Intermediate
Duration: 50:17
Speaker: : Yann LeCun and Alfredo Canziani
This module covers the concepts of gradient descent and the backpropagation algorithm and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:51:03
Speaker: : Yann LeCun
This lecture covers the concept of parameter sharing: recurrent and convolutional nets and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:59:47
Speaker: : Yann LeCun and Alfredo Canziani
This lecture covers the concept of convolutional nets in practice and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 51:40
Speaker: : Yann LeCun
This lecture discusses the concept of natural signals properties and the convolutional nets in practice and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:09:12
Speaker: : Alfredo Canziani
This lecture covers the concept of recurrent neural networks: vanilla and gated (LSTM) and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:05:36
Speaker: : Alfredo Canziani
This lecture is a foundationational lecture for the concept of energy-based models with a particular focus on the joint embedding method and latent variable energy-based models (LV-EBMs) and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:51:30
Speaker: : Yann LeCun
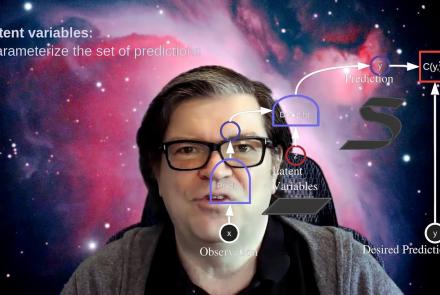
This lecture covers the concept of inference in latent variable energy based models (LV-EBMs) and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:01:04
Speaker: : Alfredo Canziani
This lecture is a foundationational lecture for the concept of energy-based models with a particular focus on the joint embedding method and latent variable energy based models (LV-EBMs) and is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:48:53
Speaker: : Yann LeCun
This tutorial covers the concept of training latent variable energy based models (LV-EBMs) and is is a part of the Deep Learning Course at NYU's Center for Data Science.
Difficulty level: Intermediate
Duration: 1:04:48
Speaker: : Alfredo Canziani
This lesson delves into the the structure of one of the brain's most elemental computational units, the neuron, and how said structure influences computational neural network models.
Difficulty level: Intermediate
Duration: 6:33
Speaker: : Marcus Ghosh
This lesson goes over the basic mechanisms of neural synapses, the space between neurons where signals may be transmitted.
Difficulty level: Intermediate
Duration: 7:03
Speaker: : Marcus Ghosh
While the previous lesson in the Neuro4ML course dealt with the mechanisms involved in individual synapses, this lesson discusses how synapses and their neurons' firing patterns may change over time.
Difficulty level: Intermediate
Duration: 4:48
Speaker: : Marcus Ghosh
This is the first of two workshops on reproducibility in science, during which participants are introduced to concepts of FAIR and open science. After discussing the definition of and need for FAIR science, participants are walked through tutorials on installing and using Github and Docker, the powerful, open-source tools for versioning and publishing code and software, respectively.
Difficulty level: Intermediate
Duration: 1:20:58
Speaker: : Erin Dickie and Sejal Patel
Topics
- Bayesian networks (2)
- Clinical neuroinformatics (2)
- Standards and Best Practices (1)
- Neuroimaging (19)
- Machine learning (9)
- Neuromorphic engineering (3)
- Tools (1)
- Animal models (1)
- Brain-hardware interfaces (1)
- Clinical neuroscience (1)
- General neuroscience (15)
- Computational neuroscience (12)
- Statistics (5)
- Computer Science (2)
- Genomics (8)
- Data science (2)
- (-) Open science (4)