This lecture presents the Medical Informatics Platform's data federation in epilepsy.
Difficulty level: Intermediate
Duration: 27:09
Speaker: : Philippe Ryvlin
This lecture aims to help researchers, students, and health care professionals understand the place for neuroinformatics in the patient journey using the exemplar of an epilepsy patient.
Difficulty level: Intermediate
Duration: 1:32:53
Speaker: : Randy Gollub & Prantik Kundu
This talk introduces data sharing initiatives in Epilepsy, particularly across Europe.
Difficulty level: Intermediate
Duration: 13:56
Speaker: : J. Helen Cross
This lesson briefly goes over the outline of the Neuroscience for Machine Learners course.
Difficulty level: Intermediate
Duration: 3:05
Speaker: : Dan Goodman
This tutorial covers the fundamentals of collaborating with Git and GitHub.
Difficulty level: Intermediate
Duration: 2:15:50
Speaker: : Elizabeth DuPre
This talk presents state-of-the-art methods for ensuring data privacy with a particular focus on medical data sharing across multiple organizations.
Difficulty level: Intermediate
Duration: 22:49
Speaker: : Barbara Carminati
This lecture talks about the usage of knowledge graphs in hospitals and related challenges of semantic interoperability.
Difficulty level: Intermediate
Duration: 24:32
Speaker: : Cristophe Gaudet-Blavignac
Course:
This lecture introduces you to the basics of the Amazon Web Services public cloud. It covers the fundamentals of cloud computing and goes through both the motivations and processes involved in moving your research computing to the cloud.
Difficulty level: Intermediate
Duration: 3:09:12
Speaker: : Amanda Tan & Ariel Rokem
This lesson continues from part one of the lecture Ontologies, Databases, and Standards, diving deeper into a description of ontologies and knowledg graphs.
Difficulty level: Intermediate
Duration: 50:18
Speaker: : Jeff Grethe
This lecture focuses on ontologies for clinical neurosciences.
Difficulty level: Intermediate
Duration: 21:54
Speaker: : Martin Hofmann-Apitius
This lesson discusses a gripping neuroscientific question: why have neurons developed the discrete action potential, or spike, as a principle method of communication?
Difficulty level: Intermediate
Duration: 9:34
Speaker: : Dan Goodman
Learn how to create a standard extracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 23:10
Speaker: : Ryan Ly
Learn how to create a standard calcium imaging dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 31:04
Speaker: : Ryan Ly
In this tutorial, you will learn how to create a standard intracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 20:23
Speaker: : Pamela Baker
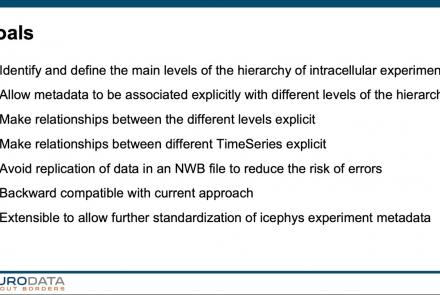
In this tutorial, you will learn how to use the icephys-metadata extension to enter meta-data detailing your experimental paradigm.
Difficulty level: Intermediate
Duration: 27:18
Speaker: : Oliver Ruebel
In this tutorial, users learn how to create a standard extracellular electrophysiology dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 45:46
Speaker: : Ben Dichter
Learn how to create a standard calcium imaging dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 39:10
Speaker: : Ben Dichter
Learn how to create a standard intracellular electrophysiology dataset in NWB.
Difficulty level: Intermediate
Duration: 20:22
Speaker: : Pamela Baker
This lesson gives an overview of the Brainstorm package for analyzing extracellular electrophysiology, including preprocessing, spike sorting, trial alignment, and spectrotemporal decomposition.
Difficulty level: Intermediate
Duration: 47:47
Speaker: : Konstantinos Nasiotis
This lesson provides an overview of the CaImAn package, as well as a demonstration of usage with NWB.
Difficulty level: Intermediate
Duration: 44:37
Speaker: : Andrea Giovannucci
Topics
- Artificial Intelligence (1)
- Provenance (1)
- EBRAINS RI (6)
- Brain-hardware interfaces (1)
- Clinical neuroscience (20)
- General neuroscience (11)
- (-) General neuroinformatics (1)
- Computational neuroscience (38)
- Statistics (2)
- (-) Computer Science (4)
- Genomics (3)
- Data science (8)
- Open science (3)
- Project management (1)
- Neuroethics (3)