In this lesson, you will learn in more detail about neuromorphic computing, that is, non-standard computational architectures that mimic some aspect of the way the brain works.
Difficulty level: Intermediate
Duration: 10:08
Speaker: : Dan Goodman
This video provides a very quick introduction to some of the neuromorphic sensing devices, and how they offer unique, low-power applications.
Difficulty level: Intermediate
Duration: 2:37
Speaker: : Dan Goodman
This lecture gives an overview of how to prepare and preprocess neuroimaging (EEG/MEG) data for use in TVB.
Difficulty level: Intermediate
Duration: 1:40:52
Speaker: : Paul Triebkorn
This lesson contains practical exercises which accompanies the first few lessons of the Neuroscience for Machine Learners (Neuro4ML) course.
Difficulty level: Intermediate
Duration: 5:58
Speaker: : Dan Goodman
This video briefly goes over the exercises accompanying Week 6 of the Neuroscience for Machine Learners (Neuro4ML) course, Understanding Neural Networks.
Difficulty level: Intermediate
Duration: 2:43
Speaker: : Marcus Ghosh
This lesson introduces the practical exercises which accompany the previous lessons on animal and human connectomes in the brain and nervous system.
Difficulty level: Intermediate
Duration: 4:10
Speaker: : Dan Goodman
This lesson discusses a gripping neuroscientific question: why have neurons developed the discrete action potential, or spike, as a principle method of communication?
Difficulty level: Intermediate
Duration: 9:34
Speaker: : Dan Goodman
Learn how to create a standard extracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 23:10
Speaker: : Ryan Ly
Learn how to create a standard calcium imaging dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 31:04
Speaker: : Ryan Ly
In this tutorial, you will learn how to create a standard intracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 20:23
Speaker: : Pamela Baker
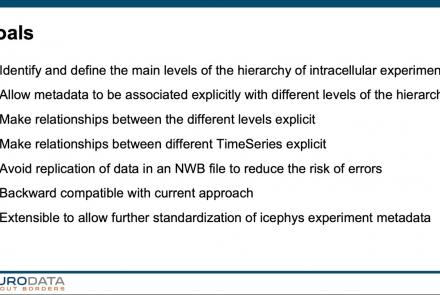
In this tutorial, you will learn how to use the icephys-metadata extension to enter meta-data detailing your experimental paradigm.
Difficulty level: Intermediate
Duration: 27:18
Speaker: : Oliver Ruebel
In this tutorial, users learn how to create a standard extracellular electrophysiology dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 45:46
Speaker: : Ben Dichter
Learn how to create a standard calcium imaging dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 39:10
Speaker: : Ben Dichter
Learn how to create a standard intracellular electrophysiology dataset in NWB.
Difficulty level: Intermediate
Duration: 20:22
Speaker: : Pamela Baker
This lesson gives an overview of the Brainstorm package for analyzing extracellular electrophysiology, including preprocessing, spike sorting, trial alignment, and spectrotemporal decomposition.
Difficulty level: Intermediate
Duration: 47:47
Speaker: : Konstantinos Nasiotis
This lesson provides an overview of the CaImAn package, as well as a demonstration of usage with NWB.
Difficulty level: Intermediate
Duration: 44:37
Speaker: : Andrea Giovannucci
This lesson gives an overview of the SpikeInterface package, including demonstration of data loading, preprocessing, spike sorting, and comparison of spike sorters.
Difficulty level: Intermediate
Duration: 1:10:28
Speaker: : Alessio Buccino
In this lesson, users will learn about the NWBWidgets package, including coverage of different data types, and information for building custom widgets within this framework.
Difficulty level: Intermediate
Duration: 47:15
Speaker: : Ben Dichter
This lesson describes the fundamentals of genomics, from central dogma to design and implementation of GWAS, to the computation, analysis, and interpretation of polygenic risk scores.
Difficulty level: Intermediate
Duration: 1:28:16
Speaker: : Dan Felsky
This is a hands-on tutorial on PLINK, the open source whole genome association analysis toolset. The aims of this tutorial are to teach users how to perform basic quality control on genetic datasets, as well as to identify and understand GWAS summary statistics.
Difficulty level: Intermediate
Duration: 1:27:18
Speaker: : Dan Felsky
Topics
- Artificial Intelligence (1)
- Notebooks (1)
- Provenance (1)
- EBRAINS RI (6)
- Animal models (1)
- Brain-hardware interfaces (1)
- Clinical neuroscience (20)
- General neuroscience (13)
- General neuroinformatics
(1)
- Computational neuroscience (52)
- Statistics (5)
- Computer Science (4)
- (-) Genomics (7)
- (-)
Data science
(10)
- Open science (5)
- Project management (1)
- Neuroethics (3)