Lesson type
Difficulty level
This lecture provides an introduction to the Brain Imaging Data Structure (BIDS), a standard for organizing human neuroimaging datasets.
Difficulty level: Intermediate
Duration: 56:49
Speaker: : Chris Gorgolewski
Course:
This lesson outlines Neurodata Without Borders (NWB), a data standard for neurophysiology which provides neuroscientists with a common standard to share, archive, use, and build analysis tools for neurophysiology data.
Difficulty level: Intermediate
Duration: 29:53
Speaker: : Oliver Ruebel
Course:
This lecture covers the rationale for developing the DAQCORD, a framework for the design, documentation, and reporting of data curation methods in order to advance the scientific rigour, reproducibility, and analysis of data.
Difficulty level: Intermediate
Duration: 17:08
Speaker: : Ari Ercole
Course:
This tutorial demonstrates how to use PyNN, a simulator-independent language for building neuronal network models, in conjunction with the neuromorphic hardware system SpiNNaker.
Difficulty level: Intermediate
Duration: 25:49
Speaker: : Christian Brenninkmeijer
Course:
This video will document the process of creating a pipeline rule for batch processing on brainlife.
Difficulty level: Intermediate
Duration: 0:57
Speaker: :
Course:
This video will document the process of launching a Jupyter Notebook for group-level analyses directly from brainlife.
Difficulty level: Intermediate
Duration: 0:53
Speaker: :
Course:
This lecture introduces you to the basics of the Amazon Web Services public cloud. It covers the fundamentals of cloud computing and goes through both the motivations and processes involved in moving your research computing to the cloud.
Difficulty level: Intermediate
Duration: 3:09:12
Speaker: : Amanda Tan & Ariel Rokem
This lesson continues from part one of the lecture Ontologies, Databases, and Standards, diving deeper into a description of ontologies and knowledg graphs.
Difficulty level: Intermediate
Duration: 50:18
Speaker: : Jeff Grethe
This lecture focuses on ontologies for clinical neurosciences.
Difficulty level: Intermediate
Duration: 21:54
Speaker: : Martin Hofmann-Apitius
Learn how to create a standard extracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 23:10
Speaker: : Ryan Ly
Learn how to create a standard calcium imaging dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 31:04
Speaker: : Ryan Ly
In this tutorial, you will learn how to create a standard intracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 20:23
Speaker: : Pamela Baker
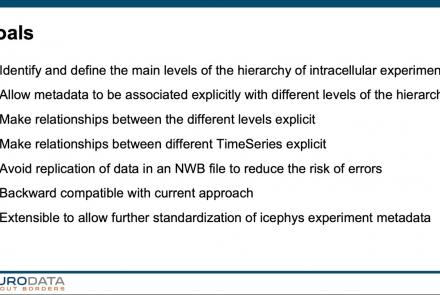
In this tutorial, you will learn how to use the icephys-metadata extension to enter meta-data detailing your experimental paradigm.
Difficulty level: Intermediate
Duration: 27:18
Speaker: : Oliver Ruebel
In this tutorial, users learn how to create a standard extracellular electrophysiology dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 45:46
Speaker: : Ben Dichter
Learn how to create a standard calcium imaging dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 39:10
Speaker: : Ben Dichter
Learn how to create a standard intracellular electrophysiology dataset in NWB.
Difficulty level: Intermediate
Duration: 20:22
Speaker: : Pamela Baker
This lesson gives an overview of the Brainstorm package for analyzing extracellular electrophysiology, including preprocessing, spike sorting, trial alignment, and spectrotemporal decomposition.
Difficulty level: Intermediate
Duration: 47:47
Speaker: : Konstantinos Nasiotis
This lesson provides an overview of the CaImAn package, as well as a demonstration of usage with NWB.
Difficulty level: Intermediate
Duration: 44:37
Speaker: : Andrea Giovannucci
This lesson gives an overview of the SpikeInterface package, including demonstration of data loading, preprocessing, spike sorting, and comparison of spike sorters.
Difficulty level: Intermediate
Duration: 1:10:28
Speaker: : Alessio Buccino
In this lesson, users will learn about the NWBWidgets package, including coverage of different data types, and information for building custom widgets within this framework.
Difficulty level: Intermediate
Duration: 47:15
Speaker: : Ben Dichter
Topics
- Artificial Intelligence (1)
- Notebooks (1)
- Provenance (1)
- EBRAINS RI (6)
- Animal models (1)
- Brain-hardware interfaces (1)
- Clinical neuroscience (20)
- General neuroscience
(16)
- General neuroinformatics (1)
- Computational neuroscience (53)
- Statistics (5)
- Computer Science (4)
- Genomics (8)
- Data science
(9)
- Open science (5)
- Project management (1)
- Neuroethics (3)