Course:
The Mouse Phenome Database (MPD) provides access to primary experimental trait data, genotypic variation, protocols and analysis tools for mouse genetic studies. Data are contributed by investigators worldwide and represent a broad scope of phenotyping endpoints and disease-related traits in naïve mice and those exposed to drugs, environmental agents or other treatments. MPD ensures rigorous curation of phenotype data and supporting documentation using relevant ontologies and controlled vocabularies. As a repository of curated and integrated data, MPD provides a means to access/re-use baseline data, as well as allows users to identify sensitized backgrounds for making new mouse models with genome editing technologies, analyze trait co-inheritance, benchmark assays in their own laboratories, and many other research applications. MPD’s primary source of funding is NIDA. For this reason, a majority of MPD data is neuro- and behavior-related.
Difficulty level: Beginner
Duration: 55:36
Speaker: : Elissa Chesler
This lecture provides an introductory overview of some of the most important concepts in software engineering.
Difficulty level: Beginner
Duration: 32:59
Speaker: : Jeff Muller
This talk enumerates the challenges regarding data accessibility and reusability inherent in the current scientific publication system, and discusses novel approaches to these challenges, such as the EBRAINS Live Papers platform.
Difficulty level: Beginner
Duration: 18:08
Speaker: : Andrew Davison
This brief video gives an introduction to the eighth session of INCF's Neuroinformatics Assembly 2023, focusing on FAIR data and the role of academic journals.
Difficulty level: Beginner
Duration: 5:57
Speaker: : Jan G. Bjaalie
This talk gives an overview of the perspectives and FAIR-aligned policies of the academic journal Public Library of Science, better known as PLOS. This journal is a nonprofit, open access publisher empowering researchers to accelerate progress in science.
Difficulty level: Beginner
Duration: 11:53
Speaker: : Iain Hrynaszkiewicz
This talk highlights a set of platform technologies, software, and data collections that close and shorten the feedback cycle in research.
Difficulty level: Beginner
Duration: 57:52
Speaker: : Satrajit Ghosh
This lesson provides an introduction to the lifecycle of EEG/ERP data, describing the various phases through which these data pass, from collection to publication.
Difficulty level: Beginner
Duration: 35:30
Speaker: : Kateřina Vařeková
In this lesson you will learn about experimental design for EEG acquisition, as well as the first phases of the EEG/ERP data lifecycle.
Difficulty level: Beginner
Duration: 30:04
Speaker: : Kateřina Vařeková
This lesson provides an overview of the current regulatory measures in place regarding experimental data security and privacy.
Difficulty level: Beginner
Duration: 31:00
Speaker: : Kateřina Vařeková
In this lesson, you will learn the appropriate methods for collection of both data and associated metadata during EEG experiments.
Difficulty level: Beginner
Duration: 29:14
Speaker: : Kateřina Vařeková
This lesson goes over methods for managing EEG/ERP data after it has been collected, from annotation to publication.
Difficulty level: Beginner
Duration: 39:25
Speaker: : Kateřina Vařeková
In this final lesson of the course, you will learn broadly about EEG signal processing, as well as specific applications which make this kind of brain signal valuable to researchers and clinicians.
Difficulty level: Beginner
Duration: 34:51
Speaker: : Kateřina Vařeková
Course:
This lesson introduces the EEGLAB toolbox, as well as motivations for its use.
Difficulty level: Beginner
Duration: 15:32
Speaker: : Arnaud Delorme
Course:
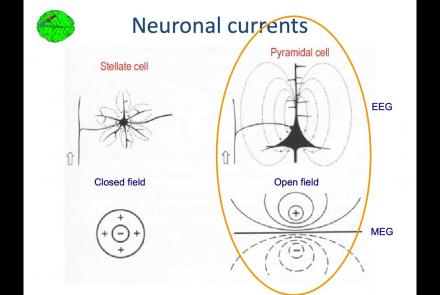
In this lesson, you will learn about the biological activity which generates and is measured by the EEG signal.
Difficulty level: Beginner
Duration: 6:53
Speaker: : Arnaud Delorme
Course:
This lesson goes over the characteristics of EEG signals when analyzed in source space (as opposed to sensor space).
Difficulty level: Beginner
Duration: 10:56
Speaker: : Arnaud Delorme
Course:
This lesson describes the development of EEGLAB as well as to what extent it is used by the research community.
Difficulty level: Beginner
Duration: 6:06
Speaker: : Arnaud Delorme
Course:
This lesson provides instruction as to how to build a processing pipeline in EEGLAB for a single participant.
Difficulty level: Beginner
Duration: 9:20
Speaker: :
Course:
Whereas the previous lesson of this course outlined how to build a processing pipeline for a single participant, this lesson discusses analysis pipelines for multiple participants simultaneously.
Difficulty level: Beginner
Duration: 10:55
Speaker: : Arnaud Delorme
Course:
In addition to outlining the motivations behind preprocessing EEG data in general, this lesson covers the first step in preprocessing data with EEGLAB, importing raw data.
Difficulty level: Beginner
Duration: 8:30
Speaker: : Arnaud Delorme
Course:
Continuing along the EEGLAB preprocessing pipeline, this tutorial walks users through how to import data events as well as EEG channel locations.
Difficulty level: Beginner
Duration: 11:53
Speaker: : Arnaud Delorme
Topics
- Artificial Intelligence (6)
- Philosophy of Science (5)
- Provenance (2)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (6)
- Assembly 2021 (29)
- Brain-hardware interfaces (13)
- Clinical neuroscience (17)
- International Brain Initiative (2)
- Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(45)
- Neuroscience (9)
- Cognitive Science (7)
- (-) Cell signaling (3)
- Brain networks (4)
- Glia (1)
- Electrophysiology (16)
- Learning and memory (3)
- Neuroanatomy (17)
- Neurobiology (7)
- Neurodegeneration (1)
- Neuroimmunology (1)
- Neural networks (4)
- (-) Neurophysiology (22)
- Neuropharmacology (2)
- (-) Synaptic plasticity (2)
- (-) Visual system (12)
- Phenome (1)
- General neuroinformatics
(15)
- Computational neuroscience (195)
- Statistics (2)
- Computer Science (15)
- Genomics (26)
- Data science
(24)
- Open science (56)
- Project management (7)
- Education (3)
- (-) Publishing (4)
- Neuroethics (37)