Course:
This lecture covers the description and characterization of an input-output relationship in a information-theoretic context.
Difficulty level: Beginner
Duration: 1:35:33
Speaker: : Jonathan D. Victor
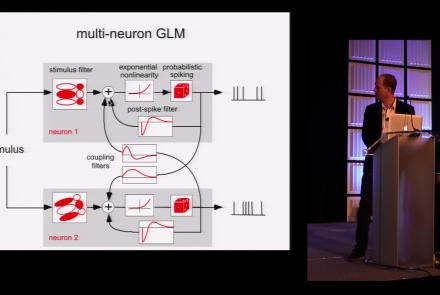
This lesson is part 1 of 2 of a tutorial on statistical models for neural data.
Difficulty level: Beginner
Duration: 1:45:48
Speaker: : Jonathan Pillow
This lesson is part 2 of 2 of a tutorial on statistical models for neural data.
Difficulty level: Beginner
Duration: 1:50:31
Speaker: : Jonathan Pillow
Course:
From the retina to the superior colliculus, the lateral geniculate nucleus into primary visual cortex and beyond, this lecture gives a tour of the mammalian visual system highlighting the Nobel-prize winning discoveries of Hubel & Wiesel.
Difficulty level: Beginner
Duration: 56:31
Speaker: : Clay Reid
Course:
From Universal Turing Machines to McCulloch-Pitts and Hopfield associative memory networks, this lecture explains what is meant by computation.
Difficulty level: Beginner
Duration: 55:27
Speaker: : Christof Koch
In this lesson you will learn about ion channels and the movement of ions across the cell membrane, one of the key mechanisms underlying neuronal communication.
Difficulty level: Beginner
Duration: 25:51
Speaker: : Carl Petersen
This lecture covers a lot of post-war developments in the science of the mind, focusing first on the cognitive revolution, and concluding with living machines.
Difficulty level: Beginner
Duration: 2:24:35
Speaker: : Paul F.M.J. Verschure
This lecture provides an overview of depression (epidemiology and course of the disorder), clinical presentation, somatic co-morbidity, and treatment options.
Difficulty level: Beginner
Duration: 37:51
Speaker: : Barbara Sperner-Unterweger
Course:
What is the difference between attention and consciousness? This lecture describes the scientific meaning of consciousness, journeys on the search for neural correlates of visual consciousness, and explores the possibility of consciousness in other beings and even non-biological structures.
Difficulty level: Beginner
Duration: 1:10:01
Speaker: : Christof Koch
Course:
This lesson provides an introduction to the myriad forms of cellular mechanisms whicn underpin healthy brain function and communication.
Difficulty level: Beginner
Duration: 12:20
Speaker: : Carl Petersen
This lesson provides an introduction to the course Cellular Mechanisms of Brain Function.
Difficulty level: Beginner
Duration: 12:20
Speaker: : Carl Petersen
This lesson presents the typical setup, equipment, and solutions used in whole-cell recording of neurons.
Difficulty level: Beginner
Duration: 09:13
Speaker: : Carl Petersen
This lesson provides an introductory overview to synaptic transmission and associated neurotransmitters.
Difficulty level: Beginner
Duration: 28:22
Speaker: : Carl Petersen
Course:
Neurodata Without Borders (NWB) is a data standard for neurophysiology that provides neuroscientists with a common standard to share, archive, use, and build common analysis tools for neurophysiology data.
Difficulty level: Beginner
Duration: 1:11
Speaker: : Ben Dichter
Course:
This lesson provides a brief introduction to the Neuroscience Information Exchange (NIX) Format data model, which allows storing fully annotated scientific datasets, i.e., data combined with rich metadata and their relations in a consistent, comprehensive format.
Difficulty level: Beginner
Duration: 1:03
Speaker: : Thomas Wachtler
Course:
In this lecture, attendees will learn how Mutant Mouse Resource and Research Center (MMRRC) archives, cryopreserves, and distributes scientifically valuable genetically engineered mouse strains and mouse ES cell lines for the genetics and biomedical research community.
Difficulty level: Beginner
Duration: 43:38
Speaker: : Kent Lloyd
This lesson provides an overview of Neurodata Without Borders (NWB), an ecosystem for neurophysiology data standardization. The lecture also introduces some NWB-enabled tools.
Difficulty level: Beginner
Duration: 29:53
Speaker: : Oliver Ruebel
This lecture discusses the FAIR principles as they apply to electrophysiology data and metadata, the building blocks for community tools and standards, platforms and grassroots initiatives, and the challenges therein.
Difficulty level: Beginner
Duration: 8:11
Speaker: : Thomas Wachtler
This lecture contains an overview of electrophysiology data reuse within the EBRAINS ecosystem.
Difficulty level: Beginner
Duration: 15:57
Speaker: : Andrew Davison
This lecture contains an overview of the Distributed Archives for Neurophysiology Data Integration (DANDI) archive, its ties to FAIR and open-source, integrations with other programs, and upcoming features.
Difficulty level: Beginner
Duration: 13:34
Speaker: : Yaroslav O. Halchenko
Topics
- Artificial Intelligence (6)
- Philosophy of Science (5)
- Provenance (2)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (6)
- Assembly 2021 (29)
- Brain-hardware interfaces (13)
- Clinical neuroscience (17)
- International Brain Initiative (2)
- Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(45)
- Neuroscience (9)
- Cognitive Science (7)
- Cell signaling (3)
- Brain networks (4)
- Glia (1)
- (-) Electrophysiology (16)
- Learning and memory (3)
- Neuroanatomy (17)
- (-) Neurobiology (7)
- Neurodegeneration (1)
- (-) Neuroimmunology (1)
- Neural networks (4)
- (-) Neurophysiology (22)
- Neuropharmacology (2)
- Synaptic plasticity (2)
- Visual system (12)
- Phenome (1)
- General neuroinformatics
(15)
- Computational neuroscience (195)
- Statistics (2)
- Computer Science (15)
- Genomics (26)
- Data science
(24)
- Open science (56)
- Project management (7)
- Education (3)
- Publishing (4)
- Neuroethics (37)