Lesson type
Difficulty level
Course:
The Mouse Phenome Database (MPD) provides access to primary experimental trait data, genotypic variation, protocols and analysis tools for mouse genetic studies. Data are contributed by investigators worldwide and represent a broad scope of phenotyping endpoints and disease-related traits in naïve mice and those exposed to drugs, environmental agents or other treatments. MPD ensures rigorous curation of phenotype data and supporting documentation using relevant ontologies and controlled vocabularies. As a repository of curated and integrated data, MPD provides a means to access/re-use baseline data, as well as allows users to identify sensitized backgrounds for making new mouse models with genome editing technologies, analyze trait co-inheritance, benchmark assays in their own laboratories, and many other research applications. MPD’s primary source of funding is NIDA. For this reason, a majority of MPD data is neuro- and behavior-related.
Difficulty level: Beginner
Duration: 55:36
Speaker: : Elissa Chesler
Course:
This lesson introduces the EEGLAB toolbox, as well as motivations for its use.
Difficulty level: Beginner
Duration: 15:32
Speaker: : Arnaud Delorme
Course:
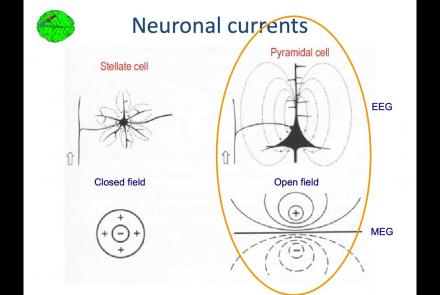
In this lesson, you will learn about the biological activity which generates and is measured by the EEG signal.
Difficulty level: Beginner
Duration: 6:53
Speaker: : Arnaud Delorme
Course:
This lesson goes over the characteristics of EEG signals when analyzed in source space (as opposed to sensor space).
Difficulty level: Beginner
Duration: 10:56
Speaker: : Arnaud Delorme
Course:
This lesson describes the development of EEGLAB as well as to what extent it is used by the research community.
Difficulty level: Beginner
Duration: 6:06
Speaker: : Arnaud Delorme
Course:
This lesson provides instruction as to how to build a processing pipeline in EEGLAB for a single participant.
Difficulty level: Beginner
Duration: 9:20
Speaker: :
Course:
Whereas the previous lesson of this course outlined how to build a processing pipeline for a single participant, this lesson discusses analysis pipelines for multiple participants simultaneously.
Difficulty level: Beginner
Duration: 10:55
Speaker: : Arnaud Delorme
Course:
In addition to outlining the motivations behind preprocessing EEG data in general, this lesson covers the first step in preprocessing data with EEGLAB, importing raw data.
Difficulty level: Beginner
Duration: 8:30
Speaker: : Arnaud Delorme
Course:
Continuing along the EEGLAB preprocessing pipeline, this tutorial walks users through how to import data events as well as EEG channel locations.
Difficulty level: Beginner
Duration: 11:53
Speaker: : Arnaud Delorme
Course:
This tutorial instructs users how to visually inspect partially pre-processed neuroimaging data in EEGLAB, specifically how to use the data browser to investigate specific channels, epochs, or events for removable artifacts, biological (e.g., eye blinks, muscle movements, heartbeat) or otherwise (e.g., corrupt channel, line noise).
Difficulty level: Beginner
Duration: 5:08
Speaker: : Arnaud Delorme
Course:
This tutorial provides instruction on how to use EEGLAB to further preprocess EEG datasets by identifying and discarding bad channels which, if left unaddressed, can corrupt and confound subsequent analysis steps.
Difficulty level: Beginner
Duration: 13:01
Speaker: : Arnaud Delorme
Course:
Users following this tutorial will learn how to identify and discard bad EEG data segments using the MATLAB toolbox EEGLAB.
Difficulty level: Beginner
Duration: 11:25
Speaker: : Arnaud Delorme
This lecture covers a wide range of aspects regarding neuroinformatics and data governance, describing both their historical developments and current trajectories. Particular tools, platforms, and standards to make your research more FAIR are also discussed.
Difficulty level: Beginner
Duration: 54:58
Speaker: : Franco Pestilli
Course:
This lesson provides an overview of GeneWeaver, a web application for the integrated cross-species analysis of functional genomics data to find convergent evidence from heterogeneous sources.
Difficulty level: Beginner
Duration: 1:03:26
Speaker: : Erich J. Baker
Course:
This lesson provides a demonstration of GeneWeaver, a system for the integration and analysis of heterogeneous functional genomics data.
Difficulty level: Beginner
Duration: 25:53
Speaker: :
Course:
This lecture outlines GeneNetwork.org, a group of linked data sets and tools used to study complex networks of genes, molecules, and higher order gene function and phenotypes.
Difficulty level: Beginner
Duration: 1:00:43
Speaker: : Robert Williams
Course:
This tutorial shows how to use the UCSC genome browser to find a list of genes in a given genomic region.
Difficulty level: Beginner
Duration: 4:32
Speaker: : UCSC Genome Browser
Course:
This tutorial shows how to find all the single nucleotide polymorphisms (SNPs) upstream from genes using the UCSC Genome Browser.
Difficulty level: Beginner
Duration: 8:13
Speaker: : UCSC Genome Browser
Course:
This tutorial demonstrates how to find all the single nucleotide polymorphisms (SNPs) in a gene using the UCSC Genome Browser.
Difficulty level: Beginner
Duration: 6:12
Speaker: : UCSC Genome Browser
Course:
The Saved Sessions feature of the Browser has been around for quite some time, but many of our users have not made full use of it. This feature offers a great way to keep track of your thinking on a particular topic.
Difficulty level: Beginner
Duration: 7:16
Speaker: : UCSC Genome Browser
Topics
- Standards and Best Practices (1)
- Machine learning (1)
- Animal models (1)
- Assembly 2021 (6)
- Clinical neuroscience (1)
- Repositories and science gateways (1)
- Resources (3)
- General neuroscience (4)
- Phenome (1)
- (-) General neuroinformatics (1)
- Computational neuroscience (20)
- Statistics (1)
- Computer Science (3)
- (-) Genomics (21)
- Data science (14)
- Open science (20)
- Education (2)
- Publishing (1)