Lesson type
Difficulty level
In this lesson, the simulation of a virtual epileptic patient is presented as an example of advanced brain simulation as a translational approach to deliver improved clinical results. You will learn about the fundamentals of epilepsy, as well as the concepts underlying epilepsy simulation. By using an iPython notebook, the detailed process of this approach is explained step by step. In the end, you are able to perform simple epilepsy simulations your own.
Difficulty level: Beginner
Duration: 1:28:53
Speaker: : Julie Courtiol
Course:
In this lesson you will learn how to simulate seizure events and epilepsy in The Virtual Brain. We will look at the paper On the Nature of Seizure Dynamics, which describes a new local model called the Epileptor, and apply this same model in The Virtual Brain. This is part 1 of 2 in a series explaining how to use the Epileptor. In this part, we focus on setting up the parameters.
Difficulty level: Beginner
Duration: 4:44
Speaker: : Paul Triebkorn
Course:
This lesson introduces the EEGLAB toolbox, as well as motivations for its use.
Difficulty level: Beginner
Duration: 15:32
Speaker: : Arnaud Delorme
Course:
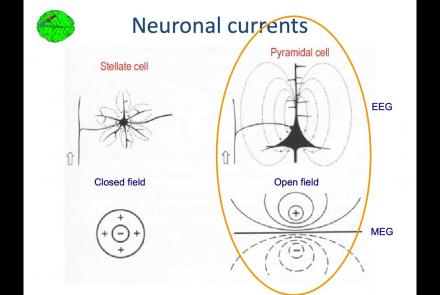
In this lesson, you will learn about the biological activity which generates and is measured by the EEG signal.
Difficulty level: Beginner
Duration: 6:53
Speaker: : Arnaud Delorme
Course:
This lesson goes over the characteristics of EEG signals when analyzed in source space (as opposed to sensor space).
Difficulty level: Beginner
Duration: 10:56
Speaker: : Arnaud Delorme
Course:
This lesson describes the development of EEGLAB as well as to what extent it is used by the research community.
Difficulty level: Beginner
Duration: 6:06
Speaker: : Arnaud Delorme
Course:
This lesson provides instruction as to how to build a processing pipeline in EEGLAB for a single participant.
Difficulty level: Beginner
Duration: 9:20
Speaker: :
Course:
Whereas the previous lesson of this course outlined how to build a processing pipeline for a single participant, this lesson discusses analysis pipelines for multiple participants simultaneously.
Difficulty level: Beginner
Duration: 10:55
Speaker: : Arnaud Delorme
Course:
In addition to outlining the motivations behind preprocessing EEG data in general, this lesson covers the first step in preprocessing data with EEGLAB, importing raw data.
Difficulty level: Beginner
Duration: 8:30
Speaker: : Arnaud Delorme
Course:
Continuing along the EEGLAB preprocessing pipeline, this tutorial walks users through how to import data events as well as EEG channel locations.
Difficulty level: Beginner
Duration: 11:53
Speaker: : Arnaud Delorme
Course:
This tutorial demonstrates how to re-reference and resample raw data in EEGLAB, why such steps are important or useful in the preprocessing pipeline, and how choices made at this step may affect subsequent analyses.
Difficulty level: Beginner
Duration: 11:48
Speaker: : Arnaud Delorme
Course:
This tutorial instructs users how to visually inspect partially pre-processed neuroimaging data in EEGLAB, specifically how to use the data browser to investigate specific channels, epochs, or events for removable artifacts, biological (e.g., eye blinks, muscle movements, heartbeat) or otherwise (e.g., corrupt channel, line noise).
Difficulty level: Beginner
Duration: 5:08
Speaker: : Arnaud Delorme
Course:
This tutorial provides instruction on how to use EEGLAB to further preprocess EEG datasets by identifying and discarding bad channels which, if left unaddressed, can corrupt and confound subsequent analysis steps.
Difficulty level: Beginner
Duration: 13:01
Speaker: : Arnaud Delorme
Course:
Users following this tutorial will learn how to identify and discard bad EEG data segments using the MATLAB toolbox EEGLAB.
Difficulty level: Beginner
Duration: 11:25
Speaker: : Arnaud Delorme
Course:
This module covers many of the types of non-invasive neurotech and neuroimaging devices including electroencephalography (EEG), electromyography (EMG), electroneurography (ENG), magnetoencephalography (MEG), and more.
Difficulty level: Beginner
Duration: 13:36
Speaker: : Harrison Canning
This opening lecture from INCF's Short Course in Neuroinformatics provides an overview of the field of neuroinformatics itself, as well as laying out an argument for the necessity for developing more sophisticated approaches towards FAIR data management principles in neuroscience.
Difficulty level: Beginner
Duration: 1:19:14
Speaker: : Maryann Martone
This lesson aims to define computational neuroscience in general terms, while providing specific examples of highly successful computational neuroscience projects.
Difficulty level: Beginner
Duration: 59:21
Speaker: : Alla Borisyuk
This lecture covers a wide range of aspects regarding neuroinformatics and data governance, describing both their historical developments and current trajectories. Particular tools, platforms, and standards to make your research more FAIR are also discussed.
Difficulty level: Beginner
Duration: 54:58
Speaker: : Franco Pestilli
Course:
Presented by the OHBM OpenScienceSIG, this lesson covers how containers can be useful for running the same software on different platforms and sharing analysis pipelines with other researchers.
Difficulty level: Beginner
Duration: 01:21:59
Speaker: : Tom Shaw & Steffen Bollmann
Serving as good refresher, this lesson explains the maths and logic concepts that are important for programmers to understand, including sets, propositional logic, conditional statements, and more.
This compilation is courtesy of freeCodeCamp.
Difficulty level: Beginner
Duration: 1:00:07
Speaker: : Shawn Grooms
Topics
- Artificial Intelligence (5)
- Philosophy of Science (5)
- Notebooks (2)
- Provenance (2)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (2)
- Assembly 2021 (28)
- Brain-hardware interfaces (13)
- Clinical neuroscience (3)
- International Brain Initiative (2)
- Repositories and science gateways (6)
- Resources (6)
- General neuroscience
(10)
- Phenome (1)
- (-)
General neuroinformatics
(4)
- Computational neuroscience (90)
- Statistics (2)
- Computer Science (8)
- Genomics (23)
- Data science
(18)
- (-) Open science (24)
- Project management (6)
- Education (2)
- Neuroethics (25)