This tutorial demonstrates how to use the image processing pipeline with the HBP collab.
Difficulty level: Beginner
Duration: 5:55
Speaker: : M. Schirner, P. Triebkorn, P. Ritter
This tutorial provides instruction on how to use the TVB-NEST toolbox co-simulation in HBP collab.
Difficulty level: Beginner
Duration: 3:11
In this tutorial, you will learn how to use TVB-NEST toolbox on your local computer.
Difficulty level: Beginner
Duration: 2:16
This tutorial provides instruction on how to perform multi-scale simulation of Alzheimer's disease on The Virtual Brain Simulation Platform.
Difficulty level: Beginner
Duration: 29:08
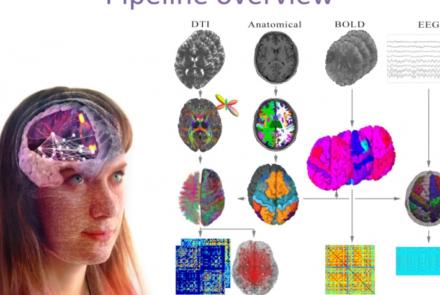
This presentation accompanies the paper entitled: An automated pipeline for constructing personalized virtual brains from multimodal neuroimaging data (see link below to download publication).
Difficulty level: Beginner
Duration: 4:56
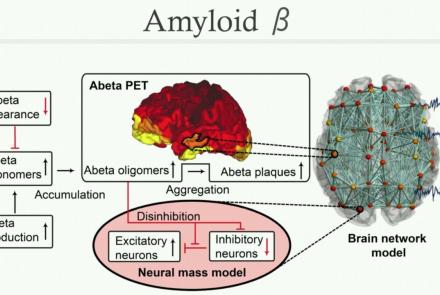
This lesson consists of a supplementary video for the publication: Inferring multi-scale neural mechanisms with brain network modelling.
Difficulty level: Beginner
Duration: 3:06
Research Resource Identifiers (RRIDs) are ID numbers assigned to help researchers cite key resources (e.g., antibodies, model organisms, and software projects) in biomedical literature to improve the transparency of research methods.
Difficulty level: Beginner
Duration: 1:01:36
Speaker: : Maryann Martone
Course:
The Mouse Phenome Database (MPD) provides access to primary experimental trait data, genotypic variation, protocols and analysis tools for mouse genetic studies. Data are contributed by investigators worldwide and represent a broad scope of phenotyping endpoints and disease-related traits in naïve mice and those exposed to drugs, environmental agents or other treatments. MPD ensures rigorous curation of phenotype data and supporting documentation using relevant ontologies and controlled vocabularies. As a repository of curated and integrated data, MPD provides a means to access/re-use baseline data, as well as allows users to identify sensitized backgrounds for making new mouse models with genome editing technologies, analyze trait co-inheritance, benchmark assays in their own laboratories, and many other research applications. MPD’s primary source of funding is NIDA. For this reason, a majority of MPD data is neuro- and behavior-related.
Difficulty level: Beginner
Duration: 55:36
Speaker: : Elissa Chesler
Course:
This lesson provides a demonstration of GeneWeaver, a system for the integration and analysis of heterogeneous functional genomics data.
Difficulty level: Beginner
Duration: 25:53
Speaker: :
Course:
This demonstration walks through how to import your data into MATLAB.
Difficulty level: Beginner
Duration: 6:10
Speaker: : MATLAB®
Course:
This lesson provides instruction regarding the various factors one must consider when preprocessing data, preparing it for statistical exploration and analyses.
Difficulty level: Beginner
Duration: 15:10
Speaker: : MATLAB®
Course:
This tutorial outlines, step by step, how to perform analysis by group and how to do change-point detection.
Difficulty level: Beginner
Duration: 2:49
Speaker: : MATLAB®
Course:
This tutorial walks through several common methods for visualizing your data in different ways depending on your data type.
Difficulty level: Beginner
Duration: 6:10
Speaker: : MATLAB®
Course:
This tutorial illustrates several ways to approach predictive modeling and machine learning with MATLAB.
Difficulty level: Beginner
Duration: 6:27
Speaker: : MATLAB®
Course:
This brief tutorial goes over how you can easily work with big data as you would with any size of data.
Difficulty level: Beginner
Duration: 3:55
Speaker: : MATLAB®
Course:
In this tutorial, you will learn how to deploy your models outside of your local MATLAB environment, enabling wider sharing and collaboration.
Difficulty level: Beginner
Duration: 3:52
Speaker: : MATLAB®
This lecture on model types introduces the advantages of modeling, provide examples of different model types, and explain what modeling is all about.
Difficulty level: Beginner
Duration: 27:48
Speaker: : Gunnar Blohm
Course:
This lecture focuses on how to get from a scientific question to a model using concrete examples. We will present a 10-step practical guide on how to succeed in modeling. This lecture contains links to 2 tutorials, lecture/tutorial slides, suggested reading list, and 3 recorded Q&A sessions.
Difficulty level: Beginner
Duration: 29:52
Speaker: : Megan Peters
Course:
This lecture focuses on the purpose of model fitting, approaches to model fitting, model fitting for linear models, and how to assess the quality and compare model fits. We will present a 10-step practical guide on how to succeed in modeling.
Difficulty level: Beginner
Duration: 26:46
Speaker: : Jan Drugowitsch
This lecture provides an overview of the generalized linear models (GLM) course, originally a part of the Neuromatch Academy (NMA), an interactive online summer school held in 2020. NMA provided participants with experiences spanning from hands-on modeling experience to meta-science interpretation skills across just about everything that could reasonably be included in the label "computational neuroscience".
Difficulty level: Beginner
Duration: 33:58
Speaker: : Cristina Savin
Topics
- Electroencephalography (EEG) (14)
- Provenance (2)
- Standards and Best Practices (1)
- Brain Medicine (1)
- Artificial Intelligence (1)
- Notebooks (1)
- Event related potential (ERP) (13)
- Digital brain atlasing (3)
- Neuroimaging (16)
- Epilepsy (1)
- Optogenetics (1)
- Neurodevelopmental disorders (1)
- Standards and best practices (1)
- Tools (13)
- Psychology (1)
- Stroke (1)
- Workflows (1)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Assembly 2021 (1)
- Brain-hardware interfaces (12)
- Repositories and science gateways (1)
- Resources (1)
- General neuroscience (5)
- Phenome (1)
- (-) Computational neuroscience (62)
- Statistics (2)
- Computer Science (3)
- Genomics (23)
- (-) Data science (12)
- (-) Open science (14)
- Education (1)
- Neuroethics (1)