Lesson type
Difficulty level
In this lesson, the simulation of a virtual epileptic patient is presented as an example of advanced brain simulation as a translational approach to deliver improved clinical results. You will learn about the fundamentals of epilepsy, as well as the concepts underlying epilepsy simulation. By using an iPython notebook, the detailed process of this approach is explained step by step. In the end, you are able to perform simple epilepsy simulations your own.
Difficulty level: Beginner
Duration: 1:28:53
Speaker: : Julie Courtiol
Course:
In this lesson you will learn how to simulate seizure events and epilepsy in The Virtual Brain. We will look at the paper On the Nature of Seizure Dynamics, which describes a new local model called the Epileptor, and apply this same model in The Virtual Brain. This is part 1 of 2 in a series explaining how to use the Epileptor. In this part, we focus on setting up the parameters.
Difficulty level: Beginner
Duration: 4:44
Speaker: : Paul Triebkorn
This talk describes the NIH-funded SPARC Data Structure, and how this project navigates ontology development while keeping in mind the FAIR science principles.
Difficulty level: Beginner
Duration: 25:44
Speaker: : Fahim Imam
Course:
This lecture covers structured data, databases, federating neuroscience-relevant databases, and ontologies.
Difficulty level: Beginner
Duration: 1:30:45
Speaker: : Maryann Martone
Course:
This lecture covers FAIR atlases, including their background and construction, as well as how they can be created in line with the FAIR principles.
Difficulty level: Beginner
Duration: 14:24
Speaker: : Heidi Kleven
This lecture covers a lot of post-war developments in the science of the mind, focusing first on the cognitive revolution, and concluding with living machines.
Difficulty level: Beginner
Duration: 2:24:35
Speaker: : Paul F.M.J. Verschure
This brief talk goes into work being done at The Alan Turing Institute to solve real-world challenges and democratize computer vision methods to support interdisciplinary and international researchers.
Difficulty level: Beginner
Duration: 7:10
Speaker: : Alden Connor & Beatriz Costa Gomes
This lesson aims to define computational neuroscience in general terms, while providing specific examples of highly successful computational neuroscience projects.
Difficulty level: Beginner
Duration: 59:21
Speaker: : Alla Borisyuk
Course:
This lecture gives an introduction to simulation, models, and the neural simulation tool NEST.
Difficulty level: Beginner
Duration: 1:48:18
Speaker: : Marc-Oliver Gewaltig
Course:
This lecture covers an Introduction to neuron anatomy and signaling, and different types of models, including the Hodgkin-Huxley model.
Difficulty level: Beginner
Duration: 1:23:01
Speaker: : Gaute Einevoll
This lecture covers an Introduction to neuron anatomy and signaling, and different types of models, including the Hodgkin-Huxley model.
Difficulty level: Beginner
Duration: 1:23:01
Speaker: : Gaute Einevoll
This lesson discuses forms of neural plasticity on many levels, including short-term, long-term, metaplasticity, and structural plasticity. During the lesson you will also be presented with examples related to the modelling of biochemical networks.
Difficulty level: Beginner
Duration: 1:11:29
Speaker: : Upi Bhalla
This lesson provides an introduction to modelling of chemical computation in the brain.
Difficulty level: Beginner
Duration: 1:00:11
Speaker: : Upi Bhalla
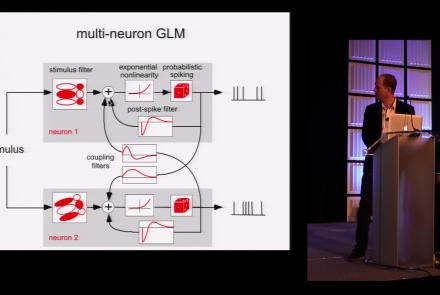
This lesson is part 1 of 2 of a tutorial on statistical models for neural data.
Difficulty level: Beginner
Duration: 1:45:48
Speaker: : Jonathan Pillow
This lesson is part 2 of 2 of a tutorial on statistical models for neural data.
Difficulty level: Beginner
Duration: 1:50:31
Speaker: : Jonathan Pillow
Course:
This lecture covers an Introduction to neuron anatomy and signaling, as well as different types of models, including the Hodgkin-Huxley model.
Difficulty level: Beginner
Duration: 1:23:01
Speaker: : Gaute Einevoll
Course:
This lecture describes forms of plasticity on many levels: short-term, long-term, metaplasticity, and structural plasticity. Included in this lecture are also examples related to modelling of biochemical networks.
Difficulty level: Beginner
Duration: 1:11:29
Speaker: : Upi Bhalla
Course:
This lesson provides an introduction to modelling of chemical computation in the brain.
Difficulty level: Beginner
Duration: 1:00:11
Speaker: : Upi Bhalla
This lesson provides an introduction to the role of models in theoretical neuroscience, particularly focusing on David Marr's work on levels of description/analysis of the brain as a complex system: computation, algorithm & representation, and implementation.
Difficulty level: Beginner
Duration: 19:26
Speaker: : Jakob Macke
In this lesson, you will learn about different types of models, model complexity, and how to choose an appropriate model.
Difficulty level: Beginner
Duration: 39:09
Speaker: : Astrid Prinz
Topics
- Artificial Intelligence (6)
- Philosophy of Science (5)
- Provenance (2)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (6)
- Assembly 2021 (29)
- Brain-hardware interfaces (13)
- Clinical neuroscience (17)
- International Brain Initiative (2)
- Repositories and science gateways (11)
- Resources (6)
- General neuroscience
(45)
- Neuroscience (9)
- Cognitive Science (7)
- Cell signaling (3)
- Brain networks (4)
- Glia (1)
- (-) Electrophysiology (16)
- Learning and memory (3)
- Neuroanatomy (17)
- Neurobiology (7)
- Neurodegeneration (1)
- Neuroimmunology (1)
- Neural networks (4)
- Neurophysiology (22)
- Neuropharmacology (2)
- Synaptic plasticity (2)
- Visual system (12)
- Phenome (1)
- General neuroinformatics
(15)
- (-) Computational neuroscience (192)
- Statistics (2)
- Computer Science (15)
- Genomics (25)
- Data science
(24)
- Open science (52)
- Project management (7)
- Education (3)
- Publishing (4)
- Neuroethics (35)