Course:
After introducing the local epileptor model in the previous two videos, we will now use it in a large-scale brain simulation. We again focus on the paper The Virtual Epileptic Patient: Individualized whole-brain models of epilepsy spread. Two simulations with different epileptogenicity across the network are visualized to show the difference in seizure spread across the cortex.
Difficulty level: Beginner
Duration: 6:36
Speaker: : Paul Triebkorn
Course:
This lecture gives an overview on the article Individual brain structure and modelling predict seizure propagation, in which 15 subjects with epilepsy were modelled to predict individual epileptogenic zones. With the TVB GUI we will model seizure spread and the effect of lesioning the connectome. The impact of cutting edges in the network on seizure spreading will be visualized.
Difficulty level: Beginner
Duration: 9:39
Speaker: : Paul Triebkorn
This lecture presents the Graphical (GUI) and Command Line (CLI) User Interface of TVB. Alongside with the speakers, explore and interact with all means necessary to generate, manipulate and visualize connectivity and network dynamics.
Difficulty level: Beginner
Duration: 1:02:16
Speaker: : Paula Popa & Mihai Andrei
This lecture briefly introduces The Virtual Brain (TVB), a multi-scale, multi-modal neuroinformatics platform for full brain network simulations using biologically realistic connectivity, as well as its potential neuroscience applications (e.g., epilepsy cases).
Difficulty level: Beginner
Duration: 8:53
Speaker: : Petra Ritter
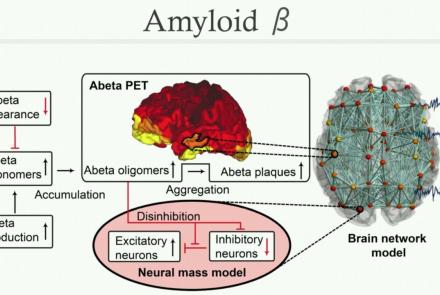
This lecture presents two recent clinical case studies using TVB: stroke recovery and dementia (due to Alzheimer’s Disease (AD)). Using a multi-scale neurophysiological model based on empirical multi-modal neuroimaging data, we show how local and global biophysical parameters characterize changes in individualized patient-specific brain dynamics, predict recovery of motor function for stroke patients, and correlate with individual differences in cognition for AD patients.
Difficulty level: Intermediate
Duration: 32:11
Speaker: : Randy McIntosh
This tutorial demonstrates how to use the image processing pipeline with the HBP collab.
Difficulty level: Beginner
Duration: 5:55
Speaker: : M. Schirner, P. Triebkorn, P. Ritter
This tutorial provides instruction on how to use the TVB-NEST toolbox co-simulation in HBP collab.
Difficulty level: Beginner
Duration: 3:11
In this tutorial, you will learn how to use TVB-NEST toolbox on your local computer.
Difficulty level: Beginner
Duration: 2:16
This tutorial provides instruction on how to perform multi-scale simulation of Alzheimer's disease on The Virtual Brain Simulation Platform.
Difficulty level: Beginner
Duration: 29:08
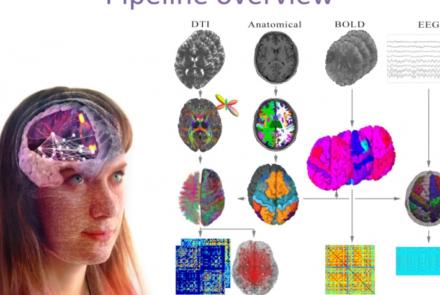
This presentation accompanies the paper entitled: An automated pipeline for constructing personalized virtual brains from multimodal neuroimaging data (see link below to download publication).
Difficulty level: Beginner
Duration: 4:56
This lesson consists of a supplementary video for the publication: Inferring multi-scale neural mechanisms with brain network modelling.
Difficulty level: Beginner
Duration: 3:06
Course:
This lesson describes the Neuroscience Gateway , which facilitates access and use of National Science Foundation High Performance Computing resources by neuroscientists.
Difficulty level: Beginner
Duration: 39:27
Speaker: : Subha Sivagnanam
Course:
This lesson introduces the EEGLAB toolbox, as well as motivations for its use.
Difficulty level: Beginner
Duration: 15:32
Speaker: : Arnaud Delorme
Course:
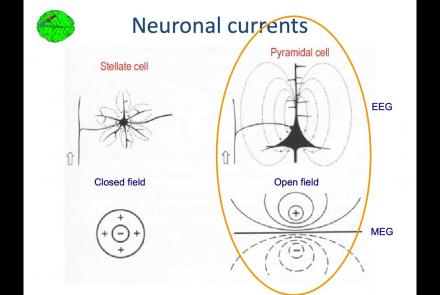
In this lesson, you will learn about the biological activity which generates and is measured by the EEG signal.
Difficulty level: Beginner
Duration: 6:53
Speaker: : Arnaud Delorme
Course:
This lesson goes over the characteristics of EEG signals when analyzed in source space (as opposed to sensor space).
Difficulty level: Beginner
Duration: 10:56
Speaker: : Arnaud Delorme
Course:
This lesson describes the development of EEGLAB as well as to what extent it is used by the research community.
Difficulty level: Beginner
Duration: 6:06
Speaker: : Arnaud Delorme
Course:
This lesson provides instruction as to how to build a processing pipeline in EEGLAB for a single participant.
Difficulty level: Beginner
Duration: 9:20
Speaker: :
Course:
Whereas the previous lesson of this course outlined how to build a processing pipeline for a single participant, this lesson discusses analysis pipelines for multiple participants simultaneously.
Difficulty level: Beginner
Duration: 10:55
Speaker: : Arnaud Delorme
Course:
In addition to outlining the motivations behind preprocessing EEG data in general, this lesson covers the first step in preprocessing data with EEGLAB, importing raw data.
Difficulty level: Beginner
Duration: 8:30
Speaker: : Arnaud Delorme
Course:
Continuing along the EEGLAB preprocessing pipeline, this tutorial walks users through how to import data events as well as EEG channel locations.
Difficulty level: Beginner
Duration: 11:53
Speaker: : Arnaud Delorme
Topics
- Standards and Best Practices (3)
- Provenance (2)
- Artificial Intelligence (1)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (2)
- Assembly 2021 (6)
- Brain-hardware interfaces (12)
- Clinical neuroscience (3)
- Repositories and science gateways (1)
- Resources (3)
- General neuroscience (17)
- Phenome (1)
- General neuroinformatics
(1)
- (-) Computational neuroscience (95)
- Statistics (5)
- Computer Science (6)
- Genomics (31)
- (-) Data science (19)
- Open science (33)
- Project management (1)
- Education (2)
- Publishing (1)
- Neuroethics (1)