Manipulate the default connectome provided with TVB to see how structural lesions effect brain dynamics. In this hands-on session you will insert lesions into the connectome within the TVB graphical user interface (GUI). Afterwards, the modified connectome will be used for simulations and the resulting activity will be analysed using functional connectivity.
Difficulty level: Beginner
Duration: 31:22
Speaker: : Paul Triebkorn
This presentation discusses the impact of data sharing in stroke.
Difficulty level: Intermediate
Duration: 16:33
Speaker: : Valeria Caso
This talks presents an overview of the potential for data federation in stroke research.
Difficulty level: Intermediate
Duration: 21:37
Speaker: : Maurizio A. Leone
Research Resource Identifiers (RRIDs) are ID numbers assigned to help researchers cite key resources (e.g., antibodies, model organisms, and software projects) in biomedical literature to improve the transparency of research methods.
Difficulty level: Beginner
Duration: 1:01:36
Speaker: : Maryann Martone
This lecture provides an overview of successful open-access projects aimed at describing complex neuroscientific models, and makes a case for expanded use of resources in support of reproducibility and validation of models against experimental data.
Difficulty level: Beginner
Duration: 1:00:39
Speaker: : Sharon Crook
This lecture provides an introduction to the Brain Imaging Data Structure (BIDS), a standard for organizing human neuroimaging datasets.
Difficulty level: Intermediate
Duration: 56:49
Speaker: : Chris Gorgolewski
This lesson provides an overview of Neurodata Without Borders (NWB), an ecosystem for neurophysiology data standardization. The lecture also introduces some NWB-enabled tools.
Difficulty level: Beginner
Duration: 29:53
Speaker: : Oliver Ruebel
Course:
This lesson outlines Neurodata Without Borders (NWB), a data standard for neurophysiology which provides neuroscientists with a common standard to share, archive, use, and build analysis tools for neurophysiology data.
Difficulty level: Intermediate
Duration: 29:53
Speaker: : Oliver Ruebel
Course:
This lecture covers the rationale for developing the DAQCORD, a framework for the design, documentation, and reporting of data curation methods in order to advance the scientific rigour, reproducibility, and analysis of data.
Difficulty level: Intermediate
Duration: 17:08
Speaker: : Ari Ercole
Course:
This tutorial demonstrates how to use PyNN, a simulator-independent language for building neuronal network models, in conjunction with the neuromorphic hardware system SpiNNaker.
Difficulty level: Intermediate
Duration: 25:49
Speaker: : Christian Brenninkmeijer
Course:
This lesson introduces the EEGLAB toolbox, as well as motivations for its use.
Difficulty level: Beginner
Duration: 15:32
Speaker: : Arnaud Delorme
Course:
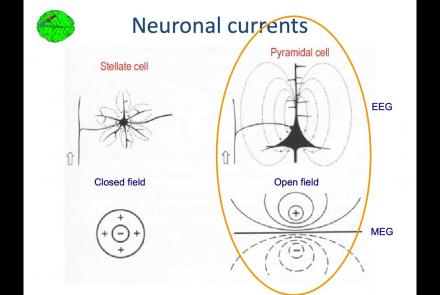
In this lesson, you will learn about the biological activity which generates and is measured by the EEG signal.
Difficulty level: Beginner
Duration: 6:53
Speaker: : Arnaud Delorme
Course:
This lesson goes over the characteristics of EEG signals when analyzed in source space (as opposed to sensor space).
Difficulty level: Beginner
Duration: 10:56
Speaker: : Arnaud Delorme
Course:
This lesson describes the development of EEGLAB as well as to what extent it is used by the research community.
Difficulty level: Beginner
Duration: 6:06
Speaker: : Arnaud Delorme
Course:
This lesson provides instruction as to how to build a processing pipeline in EEGLAB for a single participant.
Difficulty level: Beginner
Duration: 9:20
Speaker: :
Course:
Whereas the previous lesson of this course outlined how to build a processing pipeline for a single participant, this lesson discusses analysis pipelines for multiple participants simultaneously.
Difficulty level: Beginner
Duration: 10:55
Speaker: : Arnaud Delorme
Course:
In addition to outlining the motivations behind preprocessing EEG data in general, this lesson covers the first step in preprocessing data with EEGLAB, importing raw data.
Difficulty level: Beginner
Duration: 8:30
Speaker: : Arnaud Delorme
Course:
Continuing along the EEGLAB preprocessing pipeline, this tutorial walks users through how to import data events as well as EEG channel locations.
Difficulty level: Beginner
Duration: 11:53
Speaker: : Arnaud Delorme
Course:
This tutorial demonstrates how to re-reference and resample raw data in EEGLAB, why such steps are important or useful in the preprocessing pipeline, and how choices made at this step may affect subsequent analyses.
Difficulty level: Beginner
Duration: 11:48
Speaker: : Arnaud Delorme
Course:
This tutorial instructs users how to visually inspect partially pre-processed neuroimaging data in EEGLAB, specifically how to use the data browser to investigate specific channels, epochs, or events for removable artifacts, biological (e.g., eye blinks, muscle movements, heartbeat) or otherwise (e.g., corrupt channel, line noise).
Difficulty level: Beginner
Duration: 5:08
Speaker: : Arnaud Delorme
Topics
- Artificial Intelligence (6)
- Philosophy of Science (5)
- Notebooks (3)
- Provenance (3)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (3)
- Assembly 2021 (29)
- Brain-hardware interfaces (14)
- Clinical neuroscience (31)
- International Brain Initiative (2)
- Repositories and science gateways (6)
- Resources (6)
- General neuroscience
(29)
- Neuroscience (4)
- Cognitive Science (7)
- Cell signaling (4)
- Brain networks (7)
- Glia (1)
- Electrophysiology (21)
- Learning and memory (4)
- Neuroanatomy (10)
- Neurobiology (13)
- Neurodegeneration (1)
- Neuroimmunology (1)
- Neural networks (13)
- Neurophysiology (4)
- Neuropharmacology (2)
- Neuronal plasticity (16)
- Synaptic plasticity (1)
- (-) Visual system (1)
- Phenome (1)
- General neuroinformatics
(6)
- Computational neuroscience (145)
- Statistics (7)
- Computer Science (12)
- Genomics (32)
- Data science
(29)
- Open science (30)
- Project management (7)
- Education (2)
- Neuroethics (30)