Course:
This lesson describes the principles underlying functional magnetic resonance imaging (fMRI), diffusion-weighted imaging (DWI), tractography, and parcellation. These tools and concepts are explained in a broader context of neural connectivity and mental health.
Difficulty level: Intermediate
Duration: 1:47:22
Speaker: : Erin Dickie and John Griffiths
Course:
This lecture and tutorial focuses on measuring human functional brain networks, as well as how to account for inherent variability within those networks.
Difficulty level: Intermediate
Duration: 50:44
Speaker: : Caterina Gratton
Course:
This lecture presents an overview of functional brain parcellations, as well as a set of tutorials on bootstrap agregation of stable clusters (BASC) for fMRI brain parcellation.
Difficulty level: Advanced
Duration: 50:28
Speaker: : Pierre Bellec
Course:
This lecture introduces you to the basics of the Amazon Web Services public cloud. It covers the fundamentals of cloud computing and goes through both the motivations and processes involved in moving your research computing to the cloud.
Difficulty level: Intermediate
Duration: 3:09:12
Speaker: : Amanda Tan & Ariel Rokem
This lesson continues from part one of the lecture Ontologies, Databases, and Standards, diving deeper into a description of ontologies and knowledg graphs.
Difficulty level: Intermediate
Duration: 50:18
Speaker: : Jeff Grethe
This lecture focuses on ontologies for clinical neurosciences.
Difficulty level: Intermediate
Duration: 21:54
Speaker: : Martin Hofmann-Apitius
Learn how to create a standard extracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 23:10
Speaker: : Ryan Ly
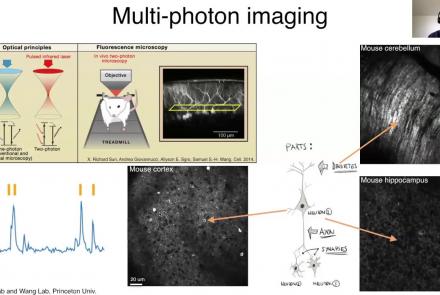
Learn how to create a standard calcium imaging dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 31:04
Speaker: : Ryan Ly
In this tutorial, you will learn how to create a standard intracellular electrophysiology dataset in NWB using Python.
Difficulty level: Intermediate
Duration: 20:23
Speaker: : Pamela Baker
In this tutorial, you will learn how to use the icephys-metadata extension to enter meta-data detailing your experimental paradigm.
Difficulty level: Intermediate
Duration: 27:18
Speaker: : Oliver Ruebel
This lesson provides instructions on how to build and share extensions in NWB.
Difficulty level: Advanced
Duration: 20:29
Speaker: : Ryan Ly
Learn how to build custom APIs for extension.
Difficulty level: Advanced
Duration: 25:40
Speaker: : Andrew Tritt
This lesson provides instruction on advanced writing strategies in HDF5 that are accessible through PyNWB.
Difficulty level: Advanced
Duration: 23:00
Speaker: : Oliver Ruebel
In this tutorial, users learn how to create a standard extracellular electrophysiology dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 45:46
Speaker: : Ben Dichter
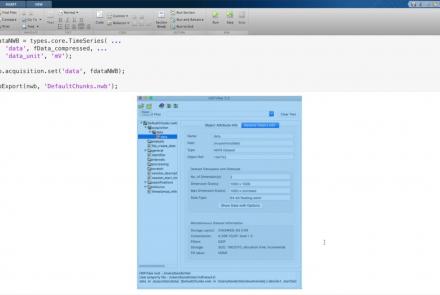
Learn how to create a standard calcium imaging dataset in NWB using MATLAB.
Difficulty level: Intermediate
Duration: 39:10
Speaker: : Ben Dichter
Learn how to create a standard intracellular electrophysiology dataset in NWB.
Difficulty level: Intermediate
Duration: 20:22
Speaker: : Pamela Baker
This lesson provides a tutorial on how to handle writing very large data in MatNWB.
Difficulty level: Advanced
Duration: 16:18
Speaker: : Ben Dichter
This lesson gives an overview of the Brainstorm package for analyzing extracellular electrophysiology, including preprocessing, spike sorting, trial alignment, and spectrotemporal decomposition.
Difficulty level: Intermediate
Duration: 47:47
Speaker: : Konstantinos Nasiotis
This lesson provides an overview of the CaImAn package, as well as a demonstration of usage with NWB.
Difficulty level: Intermediate
Duration: 44:37
Speaker: : Andrea Giovannucci
This lesson gives an overview of the SpikeInterface package, including demonstration of data loading, preprocessing, spike sorting, and comparison of spike sorters.
Difficulty level: Intermediate
Duration: 1:10:28
Speaker: : Alessio Buccino
Topics
- Artificial Intelligence (1)
- Provenance (1)
- EBRAINS RI (6)
- Brain-hardware interfaces (1)
- (-) Clinical neuroscience (20)
- General neuroscience (8)
- General neuroinformatics (1)
- Computational neuroscience (36)
- Statistics (2)
- Computer Science (2)
- Genomics (2)
- Data science (7)
- Open science (1)
- Project management (1)
- Neuroethics (3)