Course:
Overview of Day 2 of this course.
Difficulty level: Beginner
Duration: 00:03:28
Speaker: : Tristan Shuman
Course:
This talk compares various sensors and resolutions for in vivo neural recordings.
Difficulty level: Beginner
Duration: 00:24:03
Speaker: : Susie Feng, Zach Pennington, Caleb Kemere
Course:
This hands-on tutorial explains how to run your own Minion session in the MetaCell cloud using jupityr notebooks.
Difficulty level: Beginner
Duration: 01:28:03
Speaker: : Daniel Aharoni, Phil Dong
Course:
In this hands-on analysis tutorial, users will mimic a kernel crash and learn the steps to restore inputs in such a case.
Difficulty level: Beginner
Duration: 00:20:34
Speaker: : Phil Dong
Course:
This lesson will go through how to extract cells from video that has been cleaned of background noise and motion.
Difficulty level: Beginner
Duration: 01:49:40
Speaker: : Phil Dong
Course:
This final hands-on analysis tutorial walks users through the last visualization steps in the cellular data.
Difficulty level: Beginner
Duration: 00:27:23
Speaker: : Phil Dong
This lecture covers infrared LED oblique illumination for studying neuronal circuits in in vitro block-preparations of the spinal cord and brain stem.
Difficulty level: Beginner
Duration: 25:16
Speaker: : Péter Szucs
This lecture provides an introduction to the study of eye-tracking in humans.
Difficulty level: Beginner
Duration: 34:05
Speaker: : Ulrich Ettinger
This lecture covers the application of diffusion MRI for clinical and preclinical studies.
Difficulty level: Beginner
Duration: 33:10
Speaker: : Silvia de Santis
This tutorial walks participants through the application of dynamic causal modelling (DCM) to fMRI data using MATLAB. Participants are also shown various forms of DCM, how to generate and specify different models, and how to fit them to simulated neural and BOLD data.
This lesson corresponds to slides 158-187 of the PDF below.
Difficulty level: Advanced
Duration: 1:22:10
Speaker: : Peter Bedford, Povilas Karvelis
This lecture covers a lot of post-war developments in the science of the mind, focusing first on the cognitive revolution, and concluding with living machines.
Difficulty level: Beginner
Duration: 2:24:35
Speaker: : Paul F.M.J. Verschure
In this hands-on session, you will learn how to explore and work with DataLad datasets, containers, and structures using Jupyter notebooks.
Difficulty level: Beginner
Duration: 58:05
Speaker: : Michał Szczepanik
Course:
This video shows how to use the brainlife.io interface to edit the participants' info file. This file is the ParticipantInfo.json file of the Brain Imaging Data Structure (BIDS).
Difficulty level: Beginner
Duration: 0:34
Speaker: :
Course:
This quick video presents some of the various visualizers available on brainlife.io
Difficulty level: Beginner
Duration: 1:11
Speaker: :
Course:
This video demonstrates each required step for preprocessing T1w anatomical data in brainlife.io.
Difficulty level: Beginner
Duration: 3:28
Speaker: :
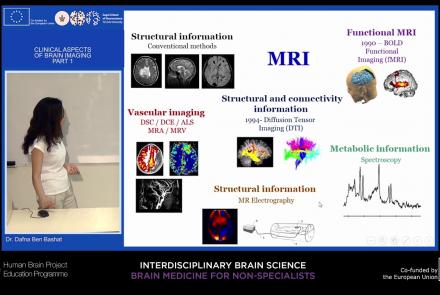
This lecture will provide an overview of neuroimaging techniques and their clinical applications.
Difficulty level: Beginner
Duration: 45:29
Speaker: : Dafna Ben Bashat
Course:
This lesson gives an introduction to simple spiking neuron models.
Difficulty level: Beginner
Duration: 48 Slides
Speaker: : Zubin Bhuyan
This lesson provides an introduction to simple spiking neuron models.
Difficulty level: Beginner
Duration: 48 Slides
Speaker: : Zubin Bhuyan
This lesson introduces the practical usage of The Virtual Brain (TVB) in its graphical user interface and via python scripts. In the graphical user interface, you are guided through its data repository, simulator, phase plane exploration tool, connectivity editor, stimulus generator, and the provided analyses. The implemented iPython notebooks of TVB are presented, and since they are public, can be used for further exploration of TVB.
Difficulty level: Beginner
Duration: 1:12:24
Speaker: : Paul Triebkorn
This hands-on tutorial focuses on a brief introduction to the GUI of TVB. You will visualize a structural connectome and use it for simulation. The local neural mass model will be explored through the phase plane viewer and a parameter space exploration will be performed to observe different dynamics of the large-scale brain model.
Difficulty level: Beginner
Duration: 23:21
Speaker: : Paul Triebkorn
Topics
- Artificial Intelligence (5)
- Philosophy of Science (5)
- Provenance (2)
- Notebooks (1)
- protein-protein interactions (1)
- Extracellular signaling (1)
- Animal models (2)
- Assembly 2021 (27)
- Brain-hardware interfaces (12)
- Clinical neuroscience (10)
- International Brain Initiative (2)
- Repositories and science gateways (5)
- Resources (6)
- General neuroscience
(10)
- Phenome (1)
- General neuroinformatics
(12)
- (-) Computational neuroscience (81)
- Statistics (2)
- Computer Science (5)
- Genomics (24)
- Data science
(17)
- Open science (23)
- Project management (3)
- Education (2)
- Neuroethics (7)