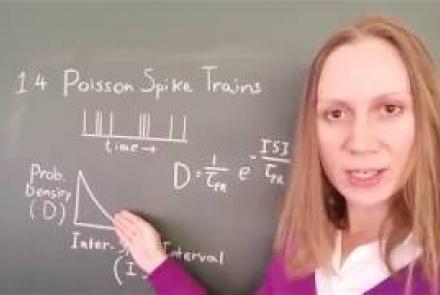This is a hands-on tutorial on PLINK, the open source whole genome association analysis toolset. The aims of this tutorial are to teach users how to perform basic quality control on genetic datasets, as well as to identify and understand GWAS summary statistics.
Difficulty level: Intermediate
Duration: 1:27:18
Speaker: : Dan Felsky
This is a tutorial on using the open-source software PRSice to calculate a set of polygenic risk scores (PRS) for a study sample. Users will also learn how to read PRS into R, visualize distributions, and perform basic association analyses.
Difficulty level: Intermediate
Duration: 1:53:34
Speaker: : Dan Felsky
This is a tutorial introducing participants to the basics of RNA-sequencing data and how to analyze its features using Seurat.
Difficulty level: Intermediate
Duration: 1:19:17
Speaker: : Sonny Chen
This tutorial demonstrates how to perform cell-type deconvolution in order to estimate how proportions of cell-types in the brain change in response to various conditions. While these techniques may be useful in addressing a wide range of scientific questions, this tutorial will focus on the cellular changes associated with major depression (MDD).
Difficulty level: Intermediate
Duration: 1:15:14
Speaker: : Keon Arbabi
Similarity Network Fusion (SNF) is a computational method for data integration across various kinds of measurements, aimed at taking advantage of the common as well as complementary information in different data types. This workshop walks participants through running SNF on EEG and genomic data using RStudio.
Difficulty level: Intermediate
Duration: 1:21:38
Speaker: : Dan Felsky
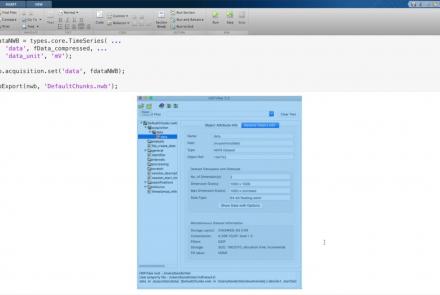
This lesson provides a tutorial on how to handle writing very large data in MatNWB.
Difficulty level: Advanced
Duration: 16:18
Speaker: : Ben Dichter
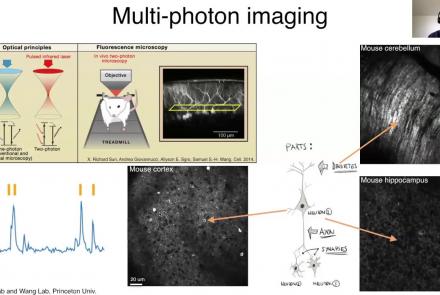
This lesson provides an overview of the CaImAn package, as well as a demonstration of usage with NWB.
Difficulty level: Intermediate
Duration: 44:37
Speaker: : Andrea Giovannucci
This lesson gives an overview of the SpikeInterface package, including demonstration of data loading, preprocessing, spike sorting, and comparison of spike sorters.
Difficulty level: Intermediate
Duration: 1:10:28
Speaker: : Alessio Buccino
In this lesson, users will learn about the NWBWidgets package, including coverage of different data types, and information for building custom widgets within this framework.
Difficulty level: Intermediate
Duration: 47:15
Speaker: : Ben Dichter
This lesson provides a brief introduction to the Computational Modeling of Neuronal Plasticity.
Difficulty level: Intermediate
Duration: 0:40
Speaker: : Florence I. Kleberg
In this lesson, you will be introducted to a type of neuronal model known as the leaky integrate-and-fire (LIF) model.
Difficulty level: Intermediate
Duration: 1:23
Speaker: : Florence I. Kleberg
This lesson goes over various potential inputs to neuronal synapses, loci of neural communication.
Difficulty level: Intermediate
Duration: 1:20
Speaker: : Florence I. Kleberg
This lesson describes the how and why behind implementing integration time steps as part of a neuronal model.
Difficulty level: Intermediate
Duration: 1:08
Speaker: : Florence I. Kleberg
In this lesson, you will learn about neural spike trains which can be characterized as having a Poisson distribution.
Difficulty level: Intermediate
Duration: 1:18
Speaker: : Florence I. Kleberg
This lesson covers spike-rate adaptation, the process by which a neuron's firing pattern decays to a low, steady-state frequency during the sustained encoding of a stimulus.
Difficulty level: Intermediate
Duration: 1:26
Speaker: : Florence I. Kleberg
This lesson provides a brief explanation of how to implement a neuron's refractory period in a computational model.
Difficulty level: Intermediate
Duration: 0:42
Speaker: : Florence I. Kleberg
In this lesson, you will learn a computational description of the process which tunes neuronal connectivity strength, spike-timing-dependent plasticity (STDP).
Difficulty level: Intermediate
Duration: 2:40
Speaker: : Florence I. Kleberg
This lesson reviews theoretical and mathematical descriptions of correlated spike trains.
Difficulty level: Intermediate
Duration: 2:54
Speaker: : Florence I. Kleberg
This lesson investigates the effect of correlated spike trains on spike-timing dependent plasticity (STDP).
Difficulty level: Intermediate
Duration: 1:43
Speaker: : Florence I. Kleberg
This lesson goes over synaptic normalisation, the homeostatic process by which groups of weighted inputs scale up or down their biases.
Difficulty level: Intermediate
Duration: 2:58
Speaker: : Florence I. Kleberg
Topics
- Bayesian networks (2)
- Cognitive neuroinformatics (1)
- Neuroimaging (15)
- Machine learning (3)
- Neuromorphic engineering (1)
- (-) Standards and best practices (4)
- Tools (1)
- Animal models (1)
- Brain-hardware interfaces (1)
- Repositories and science gateways (1)
- General neuroscience (7)
- Computational neuroscience (21)
- Statistics (4)
- (-) Computer Science (1)
- (-) Genomics (5)
- Data science (10)
- Open science (4)